Isolation and characterization of Acidithiobacillus caldus from several typical environments in China
��Դ�ڿ������ϴ�ѧѧ��(Ӣ�İ�)2007���2��
�������ߣ��ܺ鲨 ���� ���� ����� ��ǿ ��ΰ�� ������ ���»�
����ҳ�룺163 - 163
Key words��acidothermophile; biochemical traits; 16S rDNA sequence; Acidithiobacillus caldus
ժ Ҫ��
Abstract: Six strains of moderately thermophilic sulfur-oxidizing bacteria were isolated from several different typical environments in China. The identities of the isolates were confirmed by analyses of their 16S rRNA genes, and some key physiological traits. The isolates are Gram negative, rod-shaped bacteria, their optimal temperature and pH value for growth are 45-50 �� and 2.5-3.5 respectively. They are autotrophic and used elemental sulfur, sodium thiosulfate and potassium tetrathionate as electron donor, while a little glucose stimulated their growth. 16S rDNA sequences analysis reveals that the strains are phylogenetically clustered to Acidithiobacillus caldus.
������Ϣ��the National Natural Science Foundation of China
the State Basic Research Development Program of China
the China Ocean Mineral Resources Research and Development Association
J. Cent. South Univ. Technol. (2007)02-0163-07
DOI: 10.1007/s11771-007-0033-0 ![]()
Zhou Hong-bo(�ܺ鲨)1, Liu Xi(�� ��)1, Fu Bo(�� ��)1, Qiu Guan-zhou(�����)1, Huo Qiang(�� ǿ)1,
Zeng Wei-min(��ΰ��)1, Liu Jian-she(������)1, Chen Xin-hua(���»�)2
(1. School of Minerals Processing and Bioengineering, Central South University, Changsha 410083, China;
2. Key Laboratory of Marine Biogentic Resources of State Oceanic Administration, Xiamen 361005, China)
Abstract: Six strains of moderately thermophilic sulfur-oxidizing bacteria were isolated from several different typical environments in China. The identities of the isolates were confirmed by analyses of their 16S rRNA genes, and some key physiological traits. The isolates are Gram negative, rod-shaped bacteria, their optimal temperature and pH value for growth are 45-50 �� and 2.5-3.5 respectively. They are autotrophic and used elemental sulfur, sodium thiosulfate and potassium tetrathionate as electron donor, while a little glucose stimulated their growth. 16S rDNA sequences analysis reveals that the strains are phylogenetically clustered to Acidithiobacillus caldus.
Key words: acidothermophile; biochemical traits; 16S rDNA sequence; Acidithiobacillus caldus
1 Introduction
Biohydrometallurgy means the oxidation of sulphide minerals by microorganisms to extract metal[1-3]. Recovery of metals from the low-content minerals has developed into a successful technology. Acidophilic microorganisms play an important role in bioleaching system. Acidophiles utilize sulphide minerals as their energy source for growth and enhance the dissolution of minerals. In the bioleaching systems, the mostly mentioned and researched bacteria are classified as the genus Acidithiobacillus[4], including At. ferrooxidans[5], At. thiooxidans, At. albertensis and At. caldus[6]. At. ferrooxidans and At. thiooxidans are very often presented in mesophilic bioleaching environment, whereas At. caldus is one of the active strains in moderately acidothermophilic environment[7-8]. Leptospirillium species and At. caldus have been recognized as the dominant microbes in the bioreactor processing mineral ores operating at the temperature ranging from 45 �� to 55 ��[9-10].
The optimal growth temperature of At. caldus varies from 40 �� to 45 ��. In the process of bioleaching, it removed the S0 building up on the mineral surface, enhanced the contact of the microorganisms and Fe3+ to the surface of mineral and accelerated the oxidation rate[11-12].
In this paper, six moderately thermophilic sulfur-oxidizing bacteria were isolated from acid mine drainage (AMD) of mines and coal heap sites, and thermal spring water in China. A series of morphological and biochemical characterization as well as the analysis of 16S rDNA sequences were done. It was expected to provide bacterial strains to research the role of this sulfur-oxidizing moderately thermophile in the bioleaching system and to compose high efficient microflora for sulfide bioleaching.
2 Materials and methods
2.1 Isolation
Samples for microorganism isolation were taken from AMD of Dabaoshan Mineral Company in Guangdong Province, Dexing Copper Mineral Company in Jiangxi Province and coal heap sites in Changsha, Hunan Province, and hot spring water in Tengchong, Yunnan Province.
The compositions of the basal salt were as follows[13]: (NH4)2SO4 3.0 g/L, Na2SO4?10H2O 3.2 g/L, KCl 0.1 g/L, K2HPO4 0.05 g/L, MgSO4?7H2O 0.5 g/L, Ca(NO3)2 0.01 g/L. The basal salts were dissolved with 1 L distilled water and the pH value was adjusted to 3.0 with 1 mol/L sulfuric acid before being sterilized by autoclaving (121 �� for 15 min). S0 (10 g) or potassium tetrathionate (3 g) or sodium thiosulfate (10 g) was used as energy sources in the liquid media. The elemental sulfur was sterilized by autoclaving (100 �� for 30 min),repeated after 24 h for 3 times. Potassium tetrathionate and sodium thiosulfate were sterilized through a 0.22 ��m micropore filtration membrane. The media were solidified with 12 g/L agarose.
Sulfur-oxidizing microorganisms were enriched in the liquid medium containing 10 g/L S0, incubated at 50�� and shaken at 150 r/min. The enrichment cultures were transferred to the agar plates and incubated at 50 ��. After 5-7 d, many colonies formed on the surface of the agar plates. The single colonies were picked and streaked on agar plates and incubated at 50 ��. Agar plate streaks were repeated 5 times to achieve pure cultures.
2.2 Preparation of DNA
20 mL liquid cultures of each isolate in log phase were centrifuged at 2 000 r/min for 5 min to precipitate the elemental sulfur in the liquid. The supernatant was centrifuged at 10 000 r/min for 5 min to harvest the bacteria. The sedimentary cells were washed twice with de-ionized water. The DNA was extracted with DNA extraction kit (BIO BASIC INC).
2.3 Amplification of 16S rRNA gene and sequencing of polymerase chain reaction(PCR) fragments
The 16S rRNA genes of each isolates were amplified by PCR using the forward primer 63F (5��-CAG GCC TAA CAC ATG CAA GTC-3��) and reverse primer 1387R(5��-GGG CGG WGT GTA CAA GGC-3��)[12]. The PCR program was 94 �� for 8 min, followed by 32 cycles of 94 �� for 45 s, 55 �� for 45 s, and 72 �� for 90 s, and finally 72 �� for 10 min. PCR products of the expected size (approximately 1.6 kb) were excised from 10 g/L low-melting-point agarose gels and purified with the purification columns (Promega) in accordance with the manufacturer��s instructions. The PCR products were ligated to the pUCm-T vector and transformed into Escherichia coli strain DH5��. The white colonies on the Luria-Bertani (LB) plates containing ampicillin (0.1 mg/mL) and X-gal (20 mg/mL) were selected for nucleotide sequence determination.
2.4 Phylogenetic analysis
The 16S rDNA sequences of the isolates were compared with the 16S rDNA sequences deposited in the GenBank database using the BLAST program. Phylogenetic trees of the isolates and some known iron and sulfur-oxidizing acidophilic microorganisms obtained from the GenBank database were constructed by using the clustal X 1.81 software.
2.5 Physiological and biochemical characteristics
The optimal growth temperature and pH value were determined in flask batch tests. To investigate the chemolithotrophy growth characteristics, isolates were inoculated into 250 mL-flasks containing 100 mL basal salt medium and different kinds of energy sources, the flasks were inoculated with suspension cell and incubated at 45 �� and shaken at 150 r/min for 6 d.
Fermentative growth was examined in the basal salt medium containing different kinds of organic substances, the culture was incubated at 45 �� and shaken at 150 r/min.
All the above experiments were examined every 12 h with a phase-contrast, a cell counting chamber was used for cell density determination. The concentrations of ferrous iron and sulfate ion were determined every two days. Changes in ferrous ion concentrations were measured using the potassium dichromate titrimetric method[15]. Changes in SO42- were measured by using volumetric method[16]. All experiments were repeated in triplicate.
3 Results and discussion
3.1 Isolation, cellular and colonial morphology of
sulfur-oxidizing microorganisms
Six isolates D-1, D-2, T-1, J-1, S-1, S-2 were obtained from the samples taken from different locations (see Table 1), their SEM images are shown in Fig.1. The six strains are rod-shaped, Gram-negative bacteria. The cells characteristically appear singly, in pairs, or occasionally as long chains and measured 0.5 to 0.8 ��m by 1.2 to 4 ��m. The six isolates form colonies on plates with potassium tetrathionate or sodium thiosulfate as an energy source. Strains D-1 and T-1 form smooth, raised, circular and trim-edged colonies with brown center on the surface of potassium tetrathionate solid medium. Strains D-2, S-1 and J-1 form brown, coarse, flat, circular and trim-edged colonies on the surface of potassium tetrathionate solid medium. On the surface of sodium thiosulfate solid medium, strain S-2 forms white, smooth, raised, circular and trim-edged colonies.

Fig.1 SEM images of the isolates
(a) Strain D-1; (b) Strain D-2; (c) Strain T-1; (d) Strain J-1; (e) Strain S-1; (f) Strain S-2
3.2 Phylogenetic affiliationsThe 16S rRNA gene sequences of the isolates have high similarities (��99%) with the species Acidithio- bacillus caldus. The relationships of the isolates with some known iron and sulfur-oxidizing acidophilic microorganisms are shown in Fig.2. The six isolates are on the same branch with species Acidithiobacillus caldus and the isolates exhibit significant DNA homology to any other Acidithiobacillus species. The GenBank accession numbers for the nucleotide sequences and nearest relatives, based on 16S rRNA gene sequence homology of the isolates are shown in Table 1.
Table 1 GenBank accession numbers for nucleotide sequences and nearest relatives, based on 16S rRNA gene sequence homology of the isolates


Fig.2 Phylogenetic tree based on distance matrix analysis of 16S rDNA sequence
(Number in parentheses represent the sequences�� accession number in GenBank. The scale bar indicates the 0.02 evolutionar distance unit)
3.3 Effects of temperature and pH value on growth
Since the six isolates were isolated from four kinds of environments and were identified as the same species Acidithiobacillus caldus, four strains (D-1, J-1, S-1, T-1) were selected to do further physiological and metabolic characterization.
The strains D-1, J-1 and T-1 grow at temperatures varying from 15 �� to 60 ��, the yield of biomass increases with increasing temperature (from 15 �� to 45��) and their optimal growth temperature is 45 ��(Fig.3). Strain S-1 grows at temperatures varying from 15 �� to 65 ��, the yield of biomass increases with increasing temperature (from 15 �� to 50 ��) and its optimal growth temperature is 50 ��.

Fig.3 Effects of temperature on growth of isolates
All the isolates can grow in the medium with an initial pH value varying from 0.3 to 6.0 (Fig.4). Under conditions of optimal growth temperatures and different initial pH values, the cell numbers of strains D-1, S-1 and J-1 increase with pH value increasing from 1.0 to 3.5, then decease as pH value increases further, their optimal growth pH value is 3.5. The cell number of strain T-1 increases with pH value increasing from 1.0 to 2.5 and deceases when pH value increases further; it has an optimal growth at pH value 2.5.
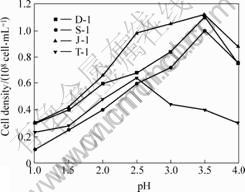
Fig.4 Effects of initial pH value on growth of isolates
3.4 Utilization of organic carbon sources
Strains D-1, J-1, S-1 and T-1 can not utilize the organic substances such as beef extract, peptone, yeast extract, glucose, fructose, sucrose, citric acid, alanine, aspartic acid, glutamic acid or glycerol as their exclusive nutrition sources. The concentrations of all the organic substances tested are 10 g/L.
Compared to their growth in medium containing elemental sulfur (10 g/L) as energy source, the growth of the four strains is inhibited when cultivated in medium containing peptone or yeast extract (Table 2), whereas the growth of the strains is stimulated by the addition of 5 g/L glucose (Table 2 and Fig.5). The cell densities of the strains growing in medium supplemented with glucose are 1.5-2.0 times higher than that in the same medium without glucose, the highest cell density is up to 9.2��108 cells/mL.
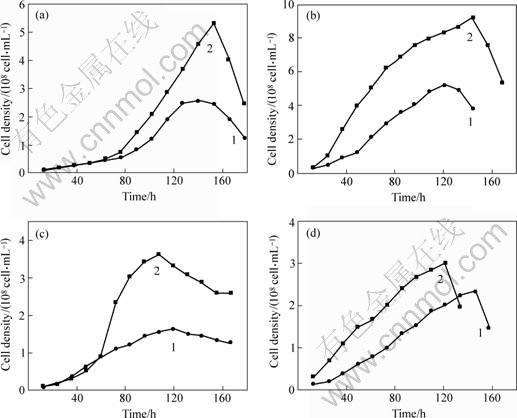
Fig.5 Effect of glucose on growth of strains D-1(a), J-1(b), S-1(c) and T-1(d) 1��S 10 g/L; 2��S 10 g/L+glucose 5 g/L
3.5 Utilization of inorganic energy sources
The four strains can not utilize ferrous sulfate, pyrite and chalcopyrite as their energy sources, while they utilize sulfur powder, potassium tetrathionate and sodium thiosulfate for growth (Table 2).
Table 2 Utilization of energy sources of isolates

The isolates are sulfur-oxidizing bacteria and produce sulfuric acid by oxidizing elemental sulfur and greatly decrease the pH value of the medium [17]. Under the conditions of their optimal growth temperatures and initial pH value of 3.0, isolates grow in liquid medium with elemental sulfur as energy source. After 8 d, the pH values of the media drop to 0.25-0.45, and the pH value of the non-inoculated medium remains the same. The productions of sulfuric acid are shown in Fig.6. The four isolates have strong ability to oxidize elemental sulfur and produce sulfuric acid. After being cultured in liquid medium for 10 d, the isolates produce 19.5-23.3 g/L SO42-, among which strain J-1 has the strongest ability to produce sulfuric acid. These data are much higher than the sulfuric acid concentration in the control test, which is only 2.5 g/L SO42-.
The isolates grow better in the medium containing S0 or K2S4O6 as the energy sources than in medium containing NaS2O3. The growth of strains D-1, J-1, S-1 and T-1 in media containing S0 or K2S4O6 is different. The results are shown in Fig.7. Since they are autotrophic bacteria growing in medium containing K2S4O6 or S0, their growth rates are much slower than that of heterotrophic bacteria. As shown in Fig.7, the growth rates of strains D-1, S-1, T-1 and J-1 in medium containing S0 are much lower than that in medium containing K2S4O6. K2S4O6 is the more available energy source for the four strains of moderately thermophilic sulfur-oxidizing bacteria.

Fig.6 Production of SO42- from elemental sulfur during strains being cultured in liquid medium containing S0

Fig.7 Growth curves of strains D-1(a), J-1(b), S-1(c) and T-1(d) in different media 1��S0 10 g/L; 2��K2S4O6 3 g/L
4 Conclusions
1) The six strains of moderately thermophilic sulfur-oxidizing bacteria isolated from four typical environments in China share the high similarity on morphology, physiological and metabolic characteristics. Their optimal temperature and pH value for growth are 45-50 �� and 2.5-3.5 respectively. They are auto- trophic and use elemental sulfur, sodium thiosulfate and potassium tetrathionate as electron donor; while a little glucose stimulates their growth.
2) 16S rDNA sequences analysis reveals that the strains are phylogenetically clustered to Acidithiobacillus caldus (more than 99% homology). It can be initially identified as Acidithiobacillus sp..
References[1] DENG Jing-shi. Bioleaching of Nickel Containing Pyrite by Moderately Thermophilic Bacteria[D]. Kunming: Kunming University of Science and Technology, 2002. (in Chinese)
[2] RAWLINGS D E. Characteristics and adaptability of iron- and sulfur-oxidizing microorganisms used for the recovery of metals from minerals and their concentrates[J]. Microbial Cell Factories, 2005, 5(4): 13-18.
[3] OLSON G J, BRIERLEY J A, BRIERLEY C L. Bioleaching review B: Processing in bioleaching: applications of microbial processes by the minerals industries[J]. Appl Microbiol Biotechnol, 2003, 63(3): 249-257.
[4] ROHWERDER T, GEHRKE T, KINZLER K, et al. Bioleaching review A: Progress in bioleaching fundamentals and mechanisms of bacterial sulfide oxidation[J]. Appl Microbiol Biotechnol, 2003, 63(3): 239-248.
[5] ZHOU Hong-bo, LIU Xi, QIU Guan-zhou, LIU Jian-she, et al. Immobilization of Acidithiobacillus ferrooxidans and ferric iron production[J]. Trans Nonferrous Met Soc China, 2006, 16(4): 931-936.
[6] KELLY D P, WOOD A P. Reclassification of some species of Thiobacillus to the newly designated genera Acidithiobacillus gen. nov., Halothiobacillus gen. nov. and Thermithiobacillus gen. nov.[J]. Syst Evol Microbiol, 2000, 50(2): 511�C516.
[7] NAOKO O, MARIEKIE G, HALLBERG B, et al. Enumeration and characterization of acidophilic microorganisms isolated from a pilot plant stirred-tank bioleaching operation[J]. Appl Environ Microbiol, 2003, 69(4): 1936-1943.
[8] FOUCHERA S, BRUNETA F B, D��HUGUESA P, et al. Evolution of the bacterial population during the batch bioleaching of a cobaltiferous pyrite in a suspended solids bubble column and comparison with a mechanically agitated reactor[J]. Hydrometallurgy, 2003, 71(1): 5�C12.
[9] DOPSON M, LINDSTROM E B. Analysis of community composition during moderately thermophilic bioleaching of pyrite, arsenical pyrite, and chalcopyrite[J]. Microbial Ecology, 2004, 48(1): 19-28.
[10] RAWLINGS D E, CORAM N J, GARDNER M N, et al. Thiobacillus caldus and Leptospirillium ferrooxians are widely distributed in continuous-flow biooxidation tanks used to treat a variety of metal-containing ores and concentrates [C]// AMILS R, BALLESTER A. Biohydrometrallurgy and the Environment Toward the Mining of the 21st Century. Part A. Amsterdam: Elsevier Press, 1999: 773-778.
[11] ROBERTSON W J, KINNUNEN P H M, PLUMB p d, et al. Moderately thermophilic ferrous-iron oxidizing bacteria isolated from a pyretic coal deposit showing spontaneous combustion[J]. Minerals Engineering, 2002, 15(11): 815-822.
[12] QIU Guan-zhou, FU Bo, ZHOU Hong-bo, et al. Isolation of a strain of Acidithiobacillus caldus and it��s role in bioleaching of chalcopyrite[J]. World J Microbiol Biotechnol, 2007, In Press.
[13] DOPSON M, B?RJE L E. Potential role of Thiobacillus caldus in arsenopyrite bioleaching[J]. Appl Environ Microbiol, 1999, 65(1): 36-40.
[14] MARCHESI J R, SATO T, WEIGHTMAN A J, et al. Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA[J]. Appl Environ Microbiol, 1998, 64(2): 795-799.
[15] RAWLINGS D E. Heavy metal mining using microbes[J]. Annual Review of Microbiology, 2002, 56: 65-91.
[16] General test method in salt industry-determination of sulfate ion, China. GB/T 13025.8-91. http://www.foodmate.net/standard/sort/3/ 4882.html
[17] SEMENZA M, VIERA M, CURUTCHET G, et al. The role of Acidithiobacillus caldus in the bioleaching of metal sulfides[J]. Latin Am Appl Research, 2002, 32(4): 303-306
Foundation item: Project (50321402) supported by the National Natural Science Foundation of China; Project(2004CB619204) supported by the State Basic
Research Development Program of China; Project (DY105-02-04-05) supported by the China Ocean Mineral Resources Research and Development Association
Received date: 2006-06-14; Accepted date: 2006-11-18
Corresponding author: ZHOU Hong-bo, Associate Proefessor, PhD; Tel: +86-731-8877216; E-mail: zhouhb@mail.csu.edu.cn
(Edited by YUAN Sai-qian)

