
Mutagenic breeding of silver-resistant Acidithiobacillus ferrooxidans and exploration of resistant mechanism
WU Xue-ling(��ѧ��), QIU Guan-zhou(�����), GAO Jian(�� ��),
DING Jian-nan(������), KANG Jian(�� ��), LIU Xin-xing(������)
Key Laboratory of Biometallurgy of Ministry of Education, School of Resources Processing and Bioengineering,
Central South University, Changsha 410083, China
Received 25 September 2006; accepted 14 December 2006
Abstract: The silver-resistant Acidithiobacillus ferrooxidans were isolated from 22 acid mine drainage (AMD) samples collected from Dexing Copper Mine and Chengmen Mountain Mine, Jiangxi Province, China. Isolate DX16 is obtained from the sample taken from Dexing Copper Mine and still carries out ferrous ion oxidation when incubated in 9K medium containing silver nitrate (240 mg/L). While isolate H1, a less resistant strain taken from Yin Mountain Mine, has a tolerate level of only 60 mg/L. Based on 16S ribosomal DNA sequencing, both bacterial 16SrDNA sequences are 100% similar to Acidithiobacillus ferrooxidans ATCC 23270. Through ultraviolet irradiation induced mutations, isolate mDX16 that is obtained from DX16 carries out ferrous ion oxidation when incubated in 9K medium containing higher concentration of silver nitrate (250 mg/L). When silver-resistant gene (SilC) analysis is carried out on the two isolates, it is seen that this gene was absent in both.
Key words: silver resistance; isolation and identification; UV mutation; Acidithiobacillus ferrooxidans; resistance mechanism
1 Introduction
Presently bioleaching is the leading mineral- processing techniques where metals are dissolved from ores into solution through oxidation and decomposing processes of the ores by microorganisms followed by the extraction of these metals from solution. Similar to all the other bioprocesses, the leaching of ores by bacteria is influenced by various environmental factors, including temperature, pH, composition of leaching medium, particle size and surrounding metal concentration. In commercial production, optimization of the above factors plays an important role in increasing leaching efficiency. At present, one of the most severe limitations in our effort to increase the efficiency of bioleaching is metal toxicity. It becomes a problem in sulphides bioleaching studies when biomining bacteria aren��t resistant to higher concentrations of metal ions. To acquire excellent bioleaching strains that grow industrially well at higher contents of metal ions, it is necessary to genetically alter bacteria via screening, adaptation and mutation. XIONG et al[1] and LONG et al[2] screened and bred Acidithiobacillus ferrooxidans (At. ferrooxidans) after ultraviolet radiation and microwave mutation and found that the oxidation activities of At .ferrooxidans increased greatly. LIU et al[3] used ores as medium, through screening, adaptation and UV mutation, and they obtained an At. ferrooxidans strain. The strain had high oxidation capacity at pH 1.2 and good adaptability when grown in uranium ores. These characteristics were expected to play a role in resource recovery and recycling of low-grade uranium ores, gangue and mineral residues. At present there are few reports about metal-resistant mutations in bacteria.
Bioleaching microorganisms that possess high resistance to metal ion toxicity, grow well and have excellent metal-leaching capability are urgently needed specially for low-grade sulphides in China. In this work, silver-resistant At. ferrooxidans were isolated and mutations were induced using UV irradiation. High silver-resistant At. ferrooxidans were screened and bred and the resistance mechanisms were investigated. The purpose of this work is to provide eugenic strains and finally to provide important theory direction for constructing metal-resistant genetically engineered bacteria.
2 Materials and methods
2.1 Recourses and enrichment of strains
22 AMD samples of water were collected from Dexing Copper Mine (labeled as DX1 to DX19, pH 1.5 to 4.5), and Chengmen Mountain Mine (labeled as H1 to H3, pH 2.0 to 4.5), Jiangxi Province, China. 9K medium [4] was used to enrich At. ferrooxidans. 5 mL of AMD samples were added to 95 mL of autoclaved 9K medium, the flasks were then placed on constant-temperature shakers at a rotation speed of 180 r/min at 30 �� until the color of the medium became reddish-brown.
2.2 Adaptation and isolation of silver-resistance strains
The concentration of silver nitrate in the 9K medium was increased gradually to adapt the silver resistance strains. 5 mL of pre-enriched culture was added to 9K medium containing 5 mg/L of silver nitrate. The culture was then incubated under the conditions described above until medium became reddish-brown. 2 mL of adapted culture was then transferred to 98 mL of fresh 9K medium containing higher concentrations (10-250 mg/L) of silver nitrate. After a series of adaptation experiments, silver-resistant colonies were isolated on 9K agar plates containing silver nitrate using the streak-plate method.
2.3 Identification of silver-resistant strains
Genomic DNA of the silver-resistant bacteria was extracted using a DNA Purification System Kit (Promega WI, USA) according to the protocol provided by the manufacturer. The 16S rDNA was amplified by Polymerase Chain Reaction. Eubacteria 16S ribosomal DNA universal primers 16SF (5��-AGAATTTGATCCTG- GCTCAG-3��) and 16SR (5��-AGAAAGGAGGTGATCC- AGCC-3��)[5] were used in this investigation. The PCR products (about 1 500 bp) were purified and 2 ��L of purified PCR products were used to clone the 16S rDNA. DNA ligation, transformation, restriction endonuclease digestion and gel electrophoresis were carried out following the protocol provided by Promiga pGEM-T Easy Vector System. The cloned products of 16S rDNA were sequenced and the sequences were compared and analyzed with the known sequences published in GenBank using the BLAST search tool of the National Centre for Biotechnology Information database.
2.4 Oxidation capacity of ferrous ions of silver- resistant strains
5 mL of culture of the silver-resistant strain at the growth logarithmic phase was added into 95 mL of fresh 9K medium containing different concentrations of silver nitrate (5-250 mg/L) and incubated in a rotary shaker (180 r/min at 30 ��). To determine the ferrous oxidation capacity of silver-resistant strain at different concentrations of silver nitrate and the highest tolerance to silver ions, the ferrous ion concentrations in the 9K medium were determined at 8 h intervals for a period of 192 h using the K2Cr2O7 titration method[6].
2.5 UV mutation of silver-resistant strains
Ultraviolet(UV) radiation was used to induce mutations in the silver-resistant isolate. The UV light was switched on and allowed to be preheated for 10 min. 10 mL of pre-cultured suspension of the adapted silver- resistant strain was transferred to plates in the logarithmic phase. A distance of 30 cm was kept between the plates and UV light. The isolates were UV irradiated for periods of 0, 30, 60, 90, 120 and 250 s, respectively at a power of 30 W. After UV irradiation, the aliquots were quickly transferred to fresh 9K medium and the flasks were placed on a rotary shaker (180 r/min at 30 ��) in the dark to prevent light-restoration. The bacterial oxidation rates of ferrous ions to ferric ions were determined from the fourth generation. Various concentrations of silver ions were added to the 9K medium, the ferrous ions oxidation rates of the mutant were determined and the silver-resistant capacities were compared.
2.6 Analysis of silver-resistance mechanism of silver- resistant strains
The SilC primer was designed according to the whole sequence of the silver-resistant SilC gene published on GenBank as follows. F: (5��-CAGCATCA- GTACCATATTC-3��), R: (5��-TGTCGGGAAGAATAG- GTAAGA-3��). Genomic DNA of the Klebsiella pneumoniae CG43 (containing SilC, presented kindly by PENG et al[7]) was extracted using a DNA Purification System Kit (Promega WI., USA) following the protocol provided by the manufacture and the SilC gene segment was amplified by PCR to determine if the primer was specific for SilC. Genomic DNA of strains DX16 and H1 was extracted using the same method above to determine if the SilC gene was present in both bacteria.
3 Results
3.1 Screening and enrichment experimental strains
In the initial enrichment experiment, 18 out of 22 samples turned the medium reddish-brown, indicating the production of jarosite, within 72 h, while the remaining samples failed to turn the medium reddish-brown after incubation for 120 h. Based on these observations, it is clear that samples 1-8, 10, 12, 14-19, H1 and H2 contains iron-oxidizing bacteria. These 18 samples were used in the subsequent studies.
3.2 Adaptation and isolation of silver-resistance strains
To screen silver resistant strain, the above 18 AMD samples were added to 9K medium respectively containing various concentrations of silver nitrate. The concentrations of silver nitrate in the 9K medium were increased gradually to adapt silver-resistant strains. When exposed to 9K medium containing 5 mg/L of silver nitrate, samples 1, 2, 8, 10, 12, 14 and 16 turned the medium reddish-brown within 72-80 h, while the process took 120 h with the remaining samples. After a series of adaptation steps, silver-resistant strains were obtained from samples DX5, DX7, DX10 and DX16 which oxidized ferrous ions to ferric ions when exposed to 220, 220, 180 and 240 mg/L of silver nitrate, respectively. As for isolate H1, the highest concentration of silver nitrate at which the organisms showed ferrous ion oxidization was 60 mg/L. Hence, strain DX16 and H1 were selected for further study due to their differences in silver resistance. DX16 and H1 were obtained using the streak-plate from 9K agar plate containing suitable concentration of silver nitrate.
3.3 Identification of strains DX16 and H1
16S rDNA fragments of DX16 and H1 were amplified, cloned and sequenced. The results show that DNA sequences of both DX16 and H1 are about 1 500 bp in size and the sequences are 100% similar to the 16S rDNA sequence of At. ferrooxidans ATCC23270 (Accession No AF465604). Therefore it is accepted that both DX16 and H1 are At. ferrooxidans.
3.4 Effect of silver on oxidation capacity of ferrous ions
In 9K medium containing 10 mg/L of silver nitrate, DX16 completely oxidized ferrous ions within 96 h but the process took 120 h when H1 was used (Fig.1). As silver nitrate concentration reached 20 mg/L, DX16 completely oxidized ferrous ions within 104 h while H1 oxidized only 55% of ferrous ions within 160 h. When DX16 was grown under the maximum silver nitrate tolerance concentration (240 mg/L), the time of complete oxidation of ferrous ions was about 192 h whereas H1 only oxidized a very small amount ferrous ions when silver nitrate concentration exceeded 60 mg/L. These results indicate that the tolerance of DX16 to silver ions is much higher than that of H1.
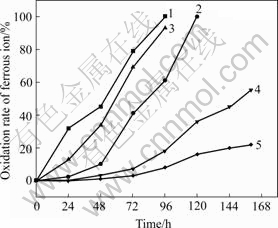
Fig.1 Comparison of oxidation rate of ferrous ion between isolate DX16 and isolate H1 in 9K medium containing silver nitrate: 1 DX16+AgNO3 (10 mg/L); 2 H1+AgNO3 (10 mg/L); 3 DX16+ AgNO3 (20 mg/L; 4 H1+AgNO3 (20 mg/L); 5 Blank+ AgNO3 (10 mg/L)
3.5 UV mutation of silver-resistant DX16
Isolate DX16 was mutated using ultraviolet radiation (30 W) at time of 0, 30, 60, 90, 120 and 250 s, then the mutants were cultured in the dark. The bacterial oxidation rates of ferrous ions to ferric ions were determined from the fourth generation and the results are shown in Fig.2.
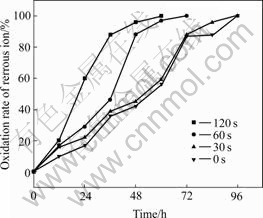
Fig.2 Oxidation rate of ferrous ion after UV mutation at different time
Fig.2 shows that the ferrous ion oxidation rates of mutants improved more than before. The ferrous ion oxidation capacity of the mutants increased slightly when the mutated-time was 30 s. Ferrous ion oxidation capacity of mutants increased obviously when the induced-time was 60 s and complete ferrous ion oxidation occurred 24 h earlier than after irradiation for 30 s. The ferrous ion oxidation capacity was highest when the strain was irradiated for 120 s taking only 60 h to completely oxidize the ferrous ions.
Figs.3 and 4 show that the highly resistant strain DX16 was mutated and the resistance of mutant mDX16 obviously increased. Before mutation, strain DX16 oxidized ferrous ions in the 9 K medium within 192 h in the presence of silver ions with a concentration of 240 mg/L. After mutation, mDX16 completely oxidized ferrous ions in the 9K medium within 170 h in the presence of silver ions with a concentration of 250 mg/L, which indicates that mDX16 has a much higher ferrous ion oxidation capacity and silver resistance than DX16.
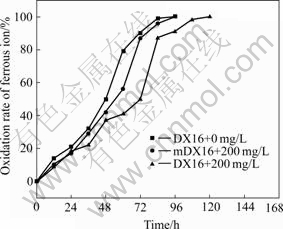
Fig.3 Comparison of oxidation rates of ferrous ion between DX16 and mDX16 in 9 K medium containing 200 mg/L silver nitrate
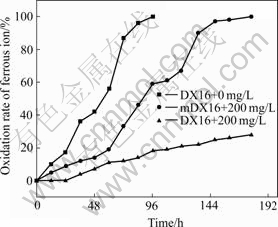
Fig.4 Comparison of oxidation rates of ferrous ion between DX16 and mDX16 in 9 K medium containing 250 mg/L silver nitrate
3.6 Analysis of silver-resistance mechanism of silver- resistant DX16
Silver-resistant gene SilC fragments were amplified by polymerase chain reaction(PCR) using self-designed SilC primer from both DX16 and H1. Klebsiella pneumoniae CG43[7] (containing SilC) was used as a positive control. The results indicated that there was a positive band in the control of Klebsiella pneumoniae CG43[7] approximately 1 500 bp in size but this band wasn��t present in DX16 and H1, which showed that the silver-resistant SilC gene was absent in both bacteria (Fig.5).
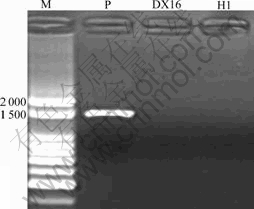
Fig.5 SilC gene agarose gel electrophoresis: M is 100 bp DNA marker; P is Klebsiella pneumoniae CG43 (containing SilC gene); DX16 and H1 are samples
4 Discussion
There are three types of metal ions that influence the activities of At. ferrooxidans. 1) Metal ions as nutrients needed for the growth of At. ferrooxidans such as Fe2+, Mg2+. 2) Metal ions that inhibit bioleaching by At. ferrooxidans such as Ni2+, Cu2+, U6+. 3) Metal ions that catalyse bioleaching by At. ferrooxidans such as Ag+, Hg2+, Bi3+, Co2+. Silver ions act as catalysts in bioleaching. They increase the solubility of copper by changing the electrochemical characters of the surface layer of chalcopyrite. QIU and WANG[8] reported that original ores taken from Dexing Copper Mine in China were bioleached with At. ferrooxidans, and the bioleaching rates of copper increased by over three times when 0.16 g of silver was added to 1 kg of ores. Furthermore, WANG et al[9] reported that the leaching rate was the highest when 10 mg/L of silver was added to 1 kg of ores of chalcopyrite. Silver ions obviously increase bioleaching rates. The catalytic capacity of silver varies with different metals. For example, silver ions have higher catalytic capability for leaching of copper than that of nickel. A possible mechanism is as follows. The covalent radius of the catalyzing ions (Ag+, Co2+, Hg2+, Bi3+) is similar to that of the leached ions (Cu2+, Ni2+, Fe2+). When bacteria destroy the crystal lattice of the surface layer of the ores, catalyzing ions entering the crystal lattice of surface layer replace the leached ions of the lattice and reacts with sulfur (catalyze-ions-sulfuration), which further destroys the crystal lattice, thereby accelerating bacterial leaching.
However, silver ions are also well known for their germicidal effect. The leaching efficiency of copper doesn��t increase any further when silver ions reach a certain concentration. Furthermore, when the silver ion concentrations increase, silver ions combine with enzymes of bacterial cells that make the cellular metabolism inactive or disordered and increase the bacterial lag phase. HU et al[10] reported that At. ferrooxidans completely oxidized the ferrous ions in 9K medium within 64 h, but the time was prolonged over 10 d in the presence of silver ions (50 mg/L); and oxidation of ferrous ion was obviously inhibited when silver ion concentration reached 100 mg/L. FAN et al[11] observed significant ferrous ion oxidation inhibition with 50 mg/L of silver nitrate.
There is a variety of metal ions in complex sulphide; eugenic bacteria with a high tolerance to toxic metals are therefore required in bioleaching. In the present study, the silver-resistance strain DX16 was obtained from AMD at Dexing Copper Mine, Jiangxi province, China through screening and adaptation. The strain DX16 grew well even in the presence of a high concentration of silver ions. DX16 completely oxidized ferrous ions in the 9K medium within 8-9 d in the presence of silver ions (240 mg/L). Whereas the low resistant isolate H1, taken from Chengmen Mountain Mine, exhibited minimal ferrous ion oxidation when silver concentration exceeded 60 mg/L, indicating that isolate DX16 has a much higher tolerance to silver than isolate H1.
To obtain eminent strains with high resistance to toxic ions used industrially, it is necessary to genetically upgrade bioleaching bacteria. At. ferrooxidans grown in acidic environment (optimize pH 2.0-3.5), are acidophilic chemolithotrophic bacteria and organic compounds strongly inhibit their growth. The majorities of mutagens play the best mutating role only in the close neutral (pH 7) environment. When pH decreases, chemical mutagens decompose into various toxic organic compounds that markedly lower their mutation effect. A physical mutagen such as ultraviolet radiation was used to induce mutations in these bacteria. XIONG et al[1] adapted At. ferrooxidans using high-sulfur gold concentration as medium induced mutations in adapted At. ferrooxidans using ultraviolet radiation and microwaves. The ferrous ion oxidation capacity of At. ferrooxidans increased by over 40-fold from 0.07 g/(L��h) to 3.18 g/(L��h). LIU et al[3] obtained a mutated At. ferrooxidans strain through screening and adaptation using ores as medium and then mutated via ultraviolet radiation. The mutated At. ferrooxidans strain had high oxidation capacity at pH 1.2 and good adaptability when growing in target uranium ores that is expected to play a role in resource recovery and recycling of low-grade uranium ores, gangue and mineral residues. XU et al[12] mutated At. ferrooxidans using microwave and investigated the bioleaching effect of low-grade chalcopyrite. The results indicated that the activities of the mutated strain increased by 39.96%, the leaching rates increased by 31.44% compared with that of the wild type and the leaching rates of easy-leaching secondary sulphide increase by 53.66%-74.97%. This shows that microwave radicalization may lead to mutations in the bioleaching bacteria and then advance bacterial activities. But up to now, it is seldom reported about metal-resistant mutants.
In the present study, through screening and adaptation, the high resistant strain DX16 was obtained and then was mutated. The resistance of the mutated strain mDX16 increases obviously. Before mutation, strain DX16 oxidized ferrous ions in 9K medium within 192 h in the presence of silver ions (240 mg/L). And the mutated strain mDX16 oxidized ferrous ions in 9K medium within 170 h in the presence of silver ions (250 mg/L), which indicates that mutant mDX16 has much higher ferrous ion oxidation capacity and silver resistance than DX16 (adapted strain) does.
In China, the majorities of the low-grade sulfuric ores contain varieties of metal ions and some of them are highly germicidal. To obtain eminent strains used industrially that have high toxic ions resistance and then to construct gene-engineering bioleaching bacteria, in this study, 16SrDNA of DX16 and H1 were sequenced and identified, and the silver-resistance mechanisms were investigated. 16S rDNA sequences were analyzed and the results showed that both isolate DX16 and isolate H1 were identified as At. ferrooxidans. At. ferrooxidans was first isolated and identified in the later 40 s from AMD[13]. The physiological and metabolic characteristics of the microorganisms have been studied widely and deeply. Presently At. ferrooxidans still play an important role in hydrometallurgy[14]. In recent years, resistance responsible genes of At. ferrooxidan have been reported such as arsenic-resistant Ars gene[15] (GenBank AF173880), mercury-resistant Mer gene [16-18] (GenBank D90110 and X57326). Silver- resistant Sil genes (GenBank, AF067954) had been already reported and sequenced in Salmonella[19], E.coli [20], Klebsiella pneumoniae[7]. Sil genes include SilCBA, SilP, SilE, SilRS etc that synthesize silver efflux protein, silver binding protein and synthesizing-regulator of silver binding protein respectively that are related to silver-resistance mechanisms[21]. However, much less information is known about the genes responsible for silver-resistance in At. ferrooxidans and these investigations also confirmed that it fails to find silver- resistant gene SilC in At. ferrooxidans. Silver-resistant mechanisms of At. ferrooxidans are still unknown.
5 Conclusions
1) The highly silver-resistant strain DX16 and low silver-resistant strain H1 were isolated from acid mine drainage samples in Dexing Copper Mine and Chengmen Mountain Mine, Jiangxi province, China. Both of the strains were identified as At. ferrooxidans.
2) Mutations were induced in the highly silver-resistant strain DX16 using UV irradiation and the mutant (mDX16) exhibits a much higher ferrous ion oxidation capacity and tolerance to silver ions.
3) Silver-resistant SilC gene analysis reveals that this gene is absent in both isolates DX16 and H1. The silver-resistance mechanisms of At. ferrooxidans require farther investigations.
Acknowledgments
The authors give special thanks to Professor CHAO Wei-liang for his valuable instruction and suggestions on this study, and to Professor PENG Hui-ling for kindly providing the bacteria Klebsiella pneumoniae CG43 used in the study.
References
[1] XIONG Ying, HU Jian-ping, LIN Bin-lan, ZHENG Cun-jiang. Study on the domestication and mutagenic selection of Thiobacillus ferrooxidans [J]. Multipurpose Utilization of Mineral Resources, 2001(6): 27-31.
[2] LONG Zhong-er, HUANG Yun-hong, CAI Zhao-ling, CONG Wei, OUYANG Fan. Adaption of Thiobacillus ferrooxidans tolerating low ph and biooxidation of ferrous sulphate [J]. The Chinese Journal of Process Engineering, 2002, 2(5): 415-419. (in Chinese)
[3] LIU Ya-jie, LI Jiang, CHEN Gong-xin, LIU Yan. Mutation breeding of Thiobacillus ferrooxidans tolerable the extremely low pH value and adaptable uranium ores [J]. Multipurpose Utilization of Mineral Resources, 2005, 4(2): 21-26.
[4] WANG Wei-ming, WANG Ying-gang, WANG Lin-fang, GAO Dan. Primary studies on the growth conditions of Thiobacillus ferrooxidans [J]. Non-Ferrous Mining and Metallurgy, 2004, 20(5): 54-56.
[5] RAINEY F A, WARD-RAINEY N, KROPPENSTEDT R M, STACKEBRANDT E. The genus nocardiopsis represents a phylogenetically coherent taxon and a distinct actinomycete lineage [J]. Proposal of Nocardiopsaceae Fam Nov International Journal of Systematic Bacteriology, 1996, 46: 1088-1092.
[6] WU Xue-ling, DING Jian-nan, GAO Jian, LIU Xin-xing, QIU Guan-zhou. Isolation and identification of metal-resistant iron- oxidizing bacteria [J]. Minerals and Metallurgical Processing, 2007, 24(1): 57-60.
[7] CHEN Y T, CHANG H Y, LAI Y C, PAN C C, TSAI S F, PENG H L. Sequencing and analysis of the large virulence plasmid pLVPK of Klebsiella pneumoniae CG43 [J]. Gene, 2004, 4(337): 189-198.
[8] QIU Guan-zhou, WANG Jun. Application of silver to catalyzing of bacterial leaching of copper ore [J]. Mining and Metallurgical Engineering, 1998, 18(3): 22-26.
[9] WANG Kang-lin, HAN Xiao-zhao, WANG Mo-hui, WANG Bai-nian. The study of catalyzing behavior of silver ions in bacterial leaching of chalcopyrite [J]. Mining and Metallurgical Engineering, 2003, 23(5): 60-62.
[10] HU Yue-hua, ZHANG Zai-hai, QIU Guan-zhou, WANG Dian-zuo. Catalystic effect of Ag+ on bacterial leaching�CA study [J]. Mining and Metallurgical Engineering, 2001, 21(1): 24-28.
[11] FAN You-jing, YANG Hong-ying, ZI Jian-wei. The influence of the ions in solution on the growth of bioleaching engineering bacteria [J]. Nonferrous Mining and Metallurgy, 2004, 20(2): 17-19.
[12] XU Xiao-jun, GONG Lei, MENG Yun-sheng, JIANG Jin-e. Bioleaching of low grade copper ores with microwave mutated T.f [J]. Nonferrous Metals, 2005, 57(2): 93-97.
[13] TEMPLE K L, COLMER A R. The autotrophic oxidation of iron by a new bacterium: Thiobacillus ferrooxidans [J]. Bacteriol, 1951(62): 605-611.
[14] SIMTH R W, MISRA M. Recent development sin the bioprocessing of minerals [J]. Mineral Processing and Extract Metallurgy Review, 1993(12): 37-60.
[15] RAWLINGS D E. The molecular genetics of Thiobacillus ferrooxidans and other mesophilic, acidophilic, chemolithotrophic, iron- or sulfur-oxidizing bacteria [J]. Hydrometallurgy, 2001(59): 187-201.
[16] SHIRATORI T, INOUE C, SUGAWARA K, KUSANO T, KITAGAWA Y. Cloning and expression of Thiobacillus ferrooxidans mercury ion resistance genes in Escherichia coli. [J]. Bacteriol, 1989(171): 3458-3464.
[17] INOUE C, SUGAWARA K, KUSANO T. Thiobacillus ferrooxidans mer operon: sequence analysis of the promoter and adjacent genes [J]. Gene, 1990, 96(1): 115-120.
[18] INOUE C, SUGAWARA K, KUSANO T. The merR regulatory gene in Thiobacillus ferrooxidans is spaced apart from the mer structural genes [J]. Mol Microbiol, 1991, 5(11): 2707-2718.
[19] GUPTA A, MATSUI K, LO J F, SILVER S. Molecular basis for resistance to silver cations in salmonella [J]. Nature Medicine, 1999, 5(2): 183-188.
[20] GUPTA A, PHUNG L T, TAYLOR D E, SILVER S. Diversity of silver resistance genes in IncH incompatibility group plasmids [J]. Microbiology, 2001, (147): 3393-3402.
[21] SILVER S. Bacterial silver resistance: molecular biology and uses and misuses of silver compounds [J]. FEMS Microbiology Reviews, 2003, (27): 341-353.
Foundation item: Project(5032142) supported by Chinese Science Foundation for Distinguished Group; Project(2004CB619201) supported by the Major State Basic Research Development Program of China; Project(50374076) supported by National Natural Science Foundation of China
Corresponding author: QIU Guan-zhou; Tel: +86-731-8879815; E-mail: xueling0714@yahoo.com.cn
(Edited by LI Xiang-qun)