Ecology detection of moderate thermophilic enrichment atLau Basin hydrothermal vents
��Դ�ڿ������ϴ�ѧѧ��(Ӣ�İ�)2011���2��
�������ߣ��ܺ鲨 ����� κ���� ����� ���»�
����ҳ�룺392 - 398
Key words��hydrothermal vents; phylogenetic analysis; enrichment culture; RCR-RFLP; microbial diversity; sediments
Abstract: Culturable thermophilic microorganisms were enriched from samples collected from Lau Basin hydrothermal vents in artificial seawater medium at 45 ��C and pH 7.0. Microbial diversities of the enriched communities were defined by performing a restriction fragment length polymorphism (RFLP) analysis of 16S rRNA gene sequences with enzymes MspI and Hin6 I. A total of 14 phylotypes have been detected by the RFLP patterns identified for 16S rRNA clone libraries of the enrichment. Analysis of sequences showed that at least four bacterial divisions presented in the clones libraries. The phyla Proteobacteria and Firmicutes were the most dominant groups. The majority of the sequences included in this analysis affiliated with Gamma Proteobacteria (71%) and Bacillus (23%). Scanning electron micrographs revealed that there were abundant rod and coccoidal forms encased in sulphur and sodium chloride precipitate. These results revealed that there were a diversity of moderate thermophilic bacterial populations thrived in Lau Basin hydrothermal vents that were previously not detected by either molecular retrieval or strain purification techniques.
J. Cent. South Univ. Technol. (2011) 18: 392-398
DOI: 10.1007/s11771-011-0709-3![]()
ZHOU Hong-bo(�ܺ鲨)1, JI Hou-guo(�����)1, WEI Man-man(κ����)1,
WANG Yu-guang(�����)1, CHEN Xin-hua(���»�)2
1. School of Minerals Processing and Bioengineering, Central South University, Changsha 410083, China;
2. Key Laboratory of Marine Biogenetic Resources, Third Institute of Oeanography,
State Ocean Administration, Xiamen 361005, China
? Central South University Press and Springer-Verlag Berlin Heidelberg 2011
Abstract: Culturable thermophilic microorganisms were enriched from samples collected from Lau Basin hydrothermal vents in artificial seawater medium at 45 ��C and pH 7.0. Microbial diversities of the enriched communities were defined by performing a restriction fragment length polymorphism (RFLP) analysis of 16S rRNA gene sequences with enzymes MspI and Hin6 I. A total of 14 phylotypes have been detected by the RFLP patterns identified for 16S rRNA clone libraries of the enrichment. Analysis of sequences showed that at least four bacterial divisions presented in the clones libraries. The phyla Proteobacteria and Firmicutes were the most dominant groups. The majority of the sequences included in this analysis affiliated with Gamma Proteobacteria (71%) and Bacillus (23%). Scanning electron micrographs revealed that there were abundant rod and coccoidal forms encased in sulphur and sodium chloride precipitate. These results revealed that there were a diversity of moderate thermophilic bacterial populations thrived in Lau Basin hydrothermal vents that were previously not detected by either molecular retrieval or strain purification techniques.
Key words: hydrothermal vents; phylogenetic analysis; enrichment culture; RCR-RFLP; microbial diversity; sediments
1 Introduction
Deep-sea hydrothermal vents show features totally unexpected until their discovery in 1970s [1]. Such extreme environments support a wide diversity of living organisms which were not thought to exist on Earth. Over the nearly three decades since the discovery of vents, physiological and microbial ecologists have attempted to understand the microbe�Cenvironment [2-5] and host�Csymbiont�Cenvironment interactions, sulfide and heavy-metal toxicity, thermal tolerances, and sensory adaptations of the vent organisms [6-8]. Many studies have revealed that thermophilic microorganisms are widespread in different hydrothermal vent environments [9-12]. However, understanding of the various thermophilic microorganisms occurring in this spatially and temporally variable environment is somewhat constrained by the relative difficulties in cultivation and isolation of them. Remarkably, the combination of traditional culture methods and molecular techniques is an efficient approach to promote the progress on the isolation and physiological and taxonomic investigations of vent thermophilic microbes. Thus, PCR-RFLP technique was chosen to identify cultivatable thermophilic microorganisms, optimize enrichment cultures and facilitate cultivation attempts. To avoid the bias brought from cultivation and PCR amplification, the microbial diversity was evaluated comprehensively. Many strategies have been employed to avoid the possible contamination during operation. All steps including sediment sampling, enrichment culture, total DNA extraction and PCR-RFLP analysis were carried out in strict sterile conditions, and many negative controls were set to ensure the reliability of PCR amplification and other experiments.
Studies on the direct isolation of microorganisms and microbial community have revealed the presence of several different physiological and phylogenetic groups of prokaryotes in the environments [10-13]. However, studies on the thermophilic microbial diversity associated with deep-sea hydrothermal vents have, for a long time, been restricted to the microscopic description of samples and the isolation of a restricted number of new thermophilic microorganisms. It is now generally accepted that the vast majority of microorganisms detectable by direct methods in marine and other natural samples, including those collected from much less extreme environments, cannot be assessed solely by traditional methods that rely on laboratory cultivation [14].
It is indicated that the chemical composition and biological community diversity near Lau Basin are distinct from the East Pacific Rise (EPR) and other well-studied vent sites in the Pacific Ocean [15]. This extreme environment is in favor of the growth of multiple mesophile microorganisms with unique physiological and metabolizing characteristic. Previous studies of culture-dependent and culture-independent surveys have revealed that mesophile microorganisms globally existed in Lau Basin hydrothermal vents sediment. However, very little is known about their taxonomic and ecological significance [16].
The goal of this study was to learn and identify the culturable thermophilic microorganisms of Lau Basin hydrothermal vent system. PCR-RFLP analysis based on 16S rRNA was performed.
2.1 Sampling
The sample collected from deep-sea sediments at Lau Basin in May, 2007, during round-the-world ocean research of Dayang Yihao (Ocean No.1) is brown- colored. The site depth is 1 772 m, and the temperature there is about 84 ��C. Immediately after sampling, they were labeled and transported into -20 ��C storage in sterile plastic tubes until they were processed.
2.2 Enrichment
In order to obtain the microorganisms in the sample as unabridged as possible, artificial seawater [17] with additional trace element as the basic medium was used, when inoculated yeast extraction (1 g/L) and peptone (1 g/L) were added as organic carbon source and FeSO4 (5 g/L) as energy source. Before incubation, the medium was autoclaved at 121 ��C for 20 min in order to obtain sterile trace elements stock solution. A sterile syringe with a 22-gauge needle was used to deliver approximately trace element from a stock solution into a sterile bottle, then the bottle was kept at -4 ��C standby.
The artificial seawater salts medium was composed of the following substances: 25.0 g/L NaCl, 0.14 g/L K2HPO4, 0.14 g/L CaCl2?2H2O, 0.25 g/L NH4Cl, 3.4 g/L MgSO4?7H2O, 4.18 g/L MgCl2?6H2O, 0.33 g/L KCl, 0.5 mg/L NiCl2?6H2O, 0.5 mg/L Na2SeO3?5H2O and 0.01 g/L Fe(NH4)2(SO4)2?6H2O.
Trace element solutions were added to a final concentration of 10 mL/L. It was composed of the followed substances: 1.5 g/L nitrilotriacetic acid, 3.0 g/L MgSO4?7H2O, 0.5 g/L MnSO4?2H2O, 1.0 g/L NaCl, 0.18 g/L CoSO4?7H2O, 0.1 g/L CaCl2?2H2O, 0.18 g/L ZnSO4?7H2O, 0.01 g/L CuSO4?5H2O, 0.02 g/L KAl(SO4)2?12H2O, 0.01 g/L H3BO3, 0.01 g/L Na2MoO4?2H2O, 0.025 g/L NiCl2?6H2O and 2 g/L NaS2O3.
Before sterilization, the pH value of the medium should be regulated to 7.0. After inoculation, the conical flasks (500 mL, filled with 200 mL sterile liquid medium) were incubated at 45 ��C. The enrichment was switched to the new medium several times before being used for the subsequent operations.
2.3 Genomic DNA extraction
After being incubated for three days, enrichments were firstly washed twice with 5 mL of 1 mol/L HCl to exclude iron oxides and then twice with sterilized distilled water. Cells were then harvested by the centrifugation and the total DNA was extracted by the methods of ZHOU et al [18] with some modifications. The protocol encompassed grinding, three cycles of freezing and thawing, chemical lysis in a high-salt extraction buffer (1.5 mol/L NaCl) by heating the suspension in the presence of sodium dodecyl sulfate (SDS) and hexadecyltrimethylammonium bromide, and a proteinase K step. The collected samples were ground with a mortar and pestle in the presence of sterile sand and liquid nitrogen, and then mixed with 13.5 mL of DNA extraction buffer (0.1 mol/L sodium phosphate [pH8.0], 0.1 mol/L EDTA, 0.1 mol/L Tris-HCl [pH 8.0], 1.5 mol/L NaCl, 1% CTAB) and 50 ��L of proteinase K (20 mg/mL) in sterile centrifuge tubes in a 37 ��C water bath for 30 min. The products were then frozen at -70 ��C and thawed by microwave heating until they boiled briefly, with a total of three times. Subsequently, 1.5 mL of 20% SDS was added, and the samples were incubated in a 65 ��C water bath for 2 h with gentle end- over-end inversions every 15-20 min. The supernatants were collected after centrifugation at 6 000g for 10 min at room temperature and transferred into 50 mL centrifuge tubes. The samples were extracted two more times by adding 4.5 mL of the extraction buffer and 0.5 mL of 20% SDS, vortexing for 10 s, incubating at 65 ��C for 10 min, and centrifuging as before. Supernatants from three cycles of extractions were combined and mixed with an equal volume of chloroformisoamyl alcohol (24:1, volume ratio). The aqueous phase was recovered by centrifugation and precipitated with 0.6-times volume of isopropanol at room temperature for 1 h. The pellet of crude nucleic acids was obtained by centrifugation at 12 000g for 20 min at room temperature, washed with cold 70% ethanol, and resuspended in 100 ��L of TE buffer (10 mmol/L Tris-HCl, 1 mmol/L Na-EDTA; pH 8.0). The obtained DNA was stored at -20 ��C and used as the template for a further PCR amplification.
2.4 PCR amplification of 16S rRNA fragments
PCR was performed on purified DNA with the universal bacterium-specific primers 27f (5'-AGA GTT TGA TCC TGG CTC AG-3') and 1492R (5'-GGT TAC CTT GTT ACG ACT T-3') [19]. Each PCR mixture (50 ��L) contained 2.5 mmol/L MgCl2, 0.2 mmol/L deoxynucleoside triphosphates, 0.2 pmol/L (each) primer, 1��PCR buffer (Fermentas), and 1U of Taq DNA polymerase (Fermentas). An initial denaturation step of 5 min at 94 ��C was followed by 32 cycles of 94 ��C for 45 s, 55 ��C for 40 s, and 72 ��C for 2 min. The final extension step was 72 ��C for 10 min. PCR amplifications were purified with E.Z.N.A Gel Extraction Kit in accordance with the manufacturer��s instructions and visualized on a 1% (mass to volume ratio) agarose gels stained with ethidium bromide.
2.5 Cloning and restriction fragment length polymorphism (RFLP) analysis
Products from the amplification reactions of expected size (about 1 500 bps) were pooled and purified before ligation.
The purified PCR products were ligated to the vector PGEM-T (Promega Corporation), and used to transform DH5a competent host cells. About 95 white colonies were randomly selected from the library. For restriction fragment length polymorphism (RFLP) and sequencing, the inserted fragments were amplified with the vectorspecific M13 primers, M13F (5'-GTAAAA- CGACGGCCAGT-3') and M13R (5'-CAGGAAAC- AGCTATGAC-3'). Each 25 ��L reaction mixture contained 1 ��L of DNA template, 2.5 mmol/L MgCl2, 0.2 mmol/L deoxynucleoside triphosphates, 0.2 pmol/L (each) primer, 1��PCR buffer (Fermentas), and 0.5 U of Taq DNA polymerase (Fermentas) (U is a unit for the amount of a particular enzyme. 1 U is defined as the amount of the enzyme that catalyzes the conversion of 1 ��mol of substrate per minute). The PCR conditions were as follows: 5 min denaturation at 94 ��C followed by 32 cycles of 45 s at 94 ��C, 40 s at 57 ��C, and 1.5 min at 72 ��C and ended with a 10 min extension at 72 ��C. The colony PCR products were digested with 2.5 U of MspI (Fermentas) and Hin6I (Fermentas) in 20 ��L reaction mixtures as recommended by the manufacturer and visualized on 3% (mass to volume ratio) agrose gels stained with ethidium bromide under UV excitation. Different banding patterns were noted and analyzed.
2.6 Phylogenetic analysis
By comparing the size and number of digested fragments, clones with unique RFLP patterns were picked out and sequenced subsequently. The sequences were firstly assembled, the vector��s fragments were discarded, and then the sequences were compared with those in the GenBank database (www.ncbi.nlm.nih.gov) by BLAST search to find closely related sequences to be used in subsequent analyses. Multiple sequence alignments of approximately 1 400 bp sequences were performed using Clustal X version1.8. The phylogenetic trees were constructed from evolutionary distances using the neighbor-joining method [20] and bootstrap analyses for 1000 replicates were performed. The phylogenetic tree was viewed using Treeview software [21].
2.7 Nucleotide sequence accession numbers
The nucleotide sequences determined in this work were deposited in the NCBI database under accession from No.GQ144342 to GQ144355.
3.1 Microscope view of enrichment medium
Light microscopic examinations revealed that these cultures consisted of rod- and coccoid-shaped cells of different sizes and types of motility. Cocci were the dominant morphotypes and cells were grouped into the particle-attached and the free-living type. The particle- attached cells were closely associated with the amorphous precipitates (Fig.1). In the enrichment cultures, cells ranged in size of 0.5-2 ��m wide and 0.5- 3 ��m long. Under the observation by electron microscopy,

Fig.1 Electron micrographs of precipitates in enrichment cultures (a) and cells closely associated with precipitates (b)
sheet-like crystal fragments could be seen at the cell surfaces and closely surrounded cells (Fig.1). X-ray diffraction analysis showed that the crystal structures were possible trace halite that was formed by the sulphur, sodium chloride and organic compounds. It was estimated that this coat could protect cells from the cruel extreme environment in which the microbes thrived (Fig.2). These crystals seemed to reveal the reason why it was very difficult to break cell wall in DNA extraction.

Fig.2 Electron micrographs of cells encased in amorphous precipitates
3.2 Phylogenetic analyses
DNA was successfully extracted (23 kb) and amplified with bacteria-specific primers from the enrichment culture. The community structure was defined in terms of the number of OTUs (unique RFLP patterns) present in a community and the abundance of individual clones within each OTU. For structure and diversity of the enrichment community, the populations were estimated by the number of OTUs present in the community. In addition, an analog of rare faction was used to check if the majority of the total diversity in the enrichment cultures had been successfully detected in the clones examined. The cumulative number of OTUs was plotted as a function of clone number (Fig.3). The bacterial 16S rRNA clones were numbered solely on the basis of initial detection, which was assumed to be stochastic [22]. After the first 95 bacterial 16S rRNA clones in DY2 clone library were examined, 14 of the 14 OTUs had been detected. Apparently, the results suggested that the clones tested in the experiment were sufficient to detect the microbial diversities and infer the level of distribution within communities of the sample.
To evaluate the phylogenetic diversity represented by the RFLP patterns identified for 16S rRNA clone libraries of the enrichments, a total of 14 16S rRNA sequences were obtained and used for phylogenetic analysis. Table 1 lists a summary of the phylotypes (determined by distance matrix and sequence similarity), the most closely related organism or clone, the similarity, and the number of clones found in each library for each phylotype based on sequencing. All sequenced clones from enrichment sample belonged to four bacterial lineages: Gamma proteobacteria, Firmicutes, Alpha proteobacteria, and Beta proteobacteria. Phylogenetic trees indicated the relationships between the sequences of clones from this study and other bacterial sequences (Fig.4). As shown in Fig.5, the distribution of clones among different bacterial divisions was uneven. In terms of absolute numbers of clones and the diversity of their sequences, Proteobacteria and Firmicutes were the most prominent groups of organisms presented in the clone libraries. Gamma Proteobacteria was the most abundant phylogenetic group within Proteobacteria, accounting for 72% of the clones in the library. Firmicutes comprised 23% of the clones in the library.

Fig.3 Distribution of genera in clone library of enrichment sample
Table 1 Summary of bacterial 16S rRNA sequences from enrichment culture DY2
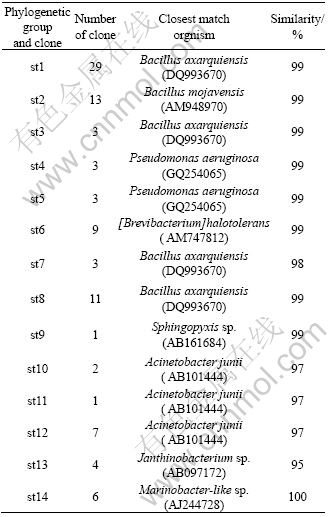

16S rRNA gene sequences of most clones were more than 97% identical to those of known bacterial species as listed in Table 1. One clone had less than 97% sequence similarity to the known bacterial species. Another clone showed 100% sequence similarity with the known Marinobacter-like sp.
These results indicated that the cultivable bacterial community in the hydrothermal vent sediment of Lau Basin is remarkably diverse.
1) Investigated cultivatable thermophilic microbial communities enriched from hydrothermal vent sediments collected at Lau Basin were determined for phylotype richness, distribution and similarity of approximately 95 16S rRNA clones. Based on 16S rRNA clone analyses, Lau Basin hydrothermal vent sediments contained a diversity range of cultivatable thermophilic microbes belonging to Proteobacteria, Firmicutes and Actinobacteria. The most dominant groups of clone libraries at Lau Basin hedrothermal vent were Proteobacteria and Firmicutes. Both Proteobacteria and Firmicutes are widespread at deep-sea hydrothermal vents. This study further expands the growing evidence that Proteobacteria and Firmicutes are important members in biogeochemical cycling of deep-sea hydrothermal ecosystems.
2) Within Proteobacteria, the members of ��, �� and �� subclasses were detected, but no ��-Proteobacteria were discovered. It is quite different from the previous in-situ investigations on the microbial communities at the hydrothermal vent sites in the Pacific Ocean and other deep-sea vent habitats, which have demonstrated the presence and dominance of bacteria belonging to the epsilon subdivision of Proteobacteria (��-Proteobacteria) that are both free-living and found in association with sediments at deep-sea hydrothermal vents. This phenomenon is possibly due to the low culturability of ��-Proteobacteria in the laboratory. The most remarkable diversity observed in the present work occurs within the gamma subdivision of Proteobacteria. The results are generally consistent with the previous reports that the subgroup of the gamma in Proteobacteria is the dominant bacterial group of the deep-sea sediment.
3) With one exception, most members of Firmicutes detected in the enrichments were grouped into the genus Bacillus. This genus is generally recognized to be among the most heterogeneous ones within the bacterial domain and has multiple genera. Thermophilic Bacillus spp. has been isolated from a wide range of thermophlic environments including deep-sea hydrothermal vents that have been recently reported. The occurrence of high-temperature, aerobic, heterotrophic spore-forming bacilli strains has been reported at Snakepit, Guaymas Basin and also Lau Basin. Additionally, thermophilic Bacillus strains growing above 45 ��C from three different active deep-sea hydrothermal vent systems (Mid-Atlantic Ridge, Guaymas Basin and Lau Basin) has been successfully isolated. The present study recovered 59 clones most closely related to two described Bacillus species, which clearly indicates that considerably diverse Bacillus populations can be readily cultured from deep-sea hydrothermal vents.
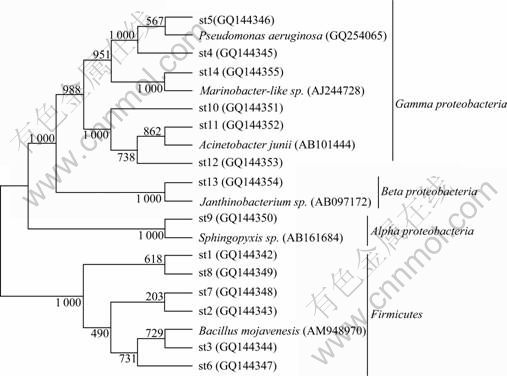
Fig.4 Phylogenetic tree based on 16s rRNA sequences from enrichment sample

Fig.5 Composition of bacterial clone libraries and percentages of clones of each phylogenetic group
4) The deep-sea hydrothermal vent was located at Lau Basin harbor, a highly diverse thermophilic prokaryotic community. Gene analysis demonstrates that Lau Basin hydrothermal vent provides a suitable habitat for cultivable thermophilic microorganisms belonging to at least four highly taxonomic lineages. These results will not only influence the choice of selective isolation procedures, but also provide further evidence that deep-sea hydrothermal vent sediments are rich sources of cultivable thermophilic microbes. And it is also suggested that it is not a lack of species diversity in deep-sea hydrothermal vent sediments that limited the ability to isolation members of novel species, but rather a need to improve selective procedures for the isolation of representative marine microbes.
5 Conclusions
1) Sheetlike crystal on the cell surface was estimated so that this coat could protect cells from the cruel extreme environment which the microbes thrived in at Lau Basin. Environment pressure including high salinity and high temperature may be the reasons that made the development.
2) Based on 16S rRNA clone analyses, the Lau Basin hydrothermal vent sediments contain a diversity range of cultivatable mesophilicthermophilic microbes belonging to the Proteobacteria, the Firmicutes, the Actinobacteria, and the Bacteroidetes. Culturable thermophilic microorganisms consist of at least four bacterial divisions. The phyla Proteobacteria and Firmicutes are the most dominant groups which are widespread at deep sea hydrothermal vents.
3) The combination of traditional culture methods and molecular techniques is an efficient approach to promote the progress on the isolation and physiological and taxonomic investigations of vent mesophilicthermophilic microbes. PCR-RFLP should be employed to make a detection of samples of which microbe diversity is unknown.
References
[1] CORLISS J B, DYMOND J, GORDON L I, EDMOND J M, VON HERZEN R P, BALLARD R D, GREEN K, WILLIAMS D, BAINBRIDGE A, CRANE K, VAN ANDEL T H. Submarine thermal springs on the Galapagos Rift [J]. Science, 1979, 203: 1073-1083.
[2] SOGIN M L, MORRISON H G., HUBER J A, WELCH D M, HUSE S M, NEAL P R, ARRIETA J M, HERNDL G J. Microbial diversity in the deep sea and the underexplored ��rare biosphere�� [J]. PNAS, 2006, 103(32): 12115-12120.
[3] EHRHARDT C J, HAYMON R M, LAMONTAGNE M G, HOLDEN P A. Evidence for hydrothermal archaea within the basaltic flanks of the East Pacific Rise [J]. Environmental Microbiology, 2007, 9(4): 900-912.
[4] PENG Xiao-tong, ZHOU Huai-yang, LI Jiang-tao, LI Ji-wei, CHEN Shun, YAO Hui-qiang, WU Zi-jun. Intracellular and extracellular mineralization of a microbial community in the Edmond deep-sea vent field environment [J]. Sedimentary Geology, 2010, 229(4): 193-206.
[5] SCHMIDT C, VUILLEMIN R, LE GALL C, GAILL F, LE BIES N. Geochemical energy sources for microbial primary production in the environment of hydrothermal vent shrimps [J]. Marine Chemistry, 2008, 108(1/2): 18-31.
[6] BATES A E, LEE R W, TUNNICLIFFE V, LAMARE M D. Deep-sea hydrothermal vent animals seek cool fluids in a highly variable thermal environment [J]. Nature Communications, 2010, 1(2): 1-6.
[7] SUZUKI Y, KOJIMA S, SASAKI T, SUZUKI M, UTSUMI T, WATANABE H, URAKAWA H, TSUCHIIDA S, NUNOURA T, HIRAYAMA H, TAKAI K, KENNETH H N, HORIKOSHI K. Host-symbiont relationships in hydrothermal vent Gastropods of the genus Alviniconcha from the southwest pacific [J]. Applied and Environmental Microbiology, 2006, 72(2): 1388-1393.
[8] BRAND G L, HORAK R V, LE BIRS N, GOFFREDI S K, CARNEY S L, GOVENAR B, YANCEY P H. Hypotaurine and thiotaurine as indicators of sulfide exposure in bivalves and vestimentiferans from hydrothermal vents and cold seeps [J]. Marine Ecology, 2007, 28(1): 208-218.
[9] MIROSHNICHENKO M L. Thermophilic microbial communities of deep-sea hydrothermal vents [J]. Microbiology, 2004, 73: 1-13.
[10] SMITH J L, CAMPBELL B J, HANSON T E, ZHANG C L, CARY C S. Nautilia profundicola sp. nov., a thermophilic, sulfur-reducing epsilonproteobacterium from deep-sea hydrothermal vents [J]. International Journal of Systematic and Evolutionary Microbiology, 2008, 58: 1598-1602.
[11] EDGCOMB V P, MOLYNEAUX S J, BOER S, WIRSEN C O, SAITO M, ATKINS M S, LLOYD K, TESKE A. Survival and growth of two heterotrophic hydrothermal vent archaea, Pyrococcus strain GB-D and Thermococcus fumicolans, under low pH and high sulfide concentrations in combination with high temperature and pressure regimes [J]. Extremophiles, 2007, 11(2): 329-342.
[12] ALAIN K, POSTEC A, GRINSARD E, LESONGEUR F, PRIEUR D, GODFROY A. Thermodesulfatator atlanticus sp. nov., a thermophilic, chemolithoautotrophic, sulfate-reducing bacterium isolated from a Mid-Atlantic Ridge hydrothermal vent [J]. International Journal of Systematic and Evolutionary Microbiology, 2010, 60: 33-38.
[13] WARD D M, BATESON M M, WELLER R, RUFF-ROBERTS A L. Ribosomal RNA analysis of microorganisms as they occur in nature [J]. Advances in Microbial Ecology, 1992, 12: 219-286.
[14] EDWARDS K J, BACH W, MCCOLLOM T M, ROGERS D R. Neutrophilic iron-oxidizing bacteria in the ocean: their habitats, diversity, and roles in mineral deposition, rock alteration, and biomass production in the deep-sea [J]. Geomicrobiology Journal, 2004, 21: 393-404.
[15] MICHAEL P, SEEWALD J. Focus on: Studies at the Lau Basin [J]. Ridge2000 Events, 2007, 2(1): 11-21.
[16] BURGAUD G, LE CALVEZ T, ARZUR D, VANDENKOORNHUYSE P, BARBIER G. Diversity of culturable marine filamentous fungi from deep-sea hydrothermal vents [J]. Environmental Microbiology, 2009, 11(6): 1588-1600.
[17] AKO Y, TAKAI K, ISHIDA Y, ARITSUNE S T, KATAYAMA Y. Rhodothermus obamensis sp. nov., a modern lineage of extremely thermophilic marine bacteria [J]. International Journal of Systematic Bacteriology, 1996, 46: 1099-1104.
[18] ZHOU J Z, BRUNS M A, TIEDJE J M. DNA recovery from soils of diverse composition [J]. Applied and Environmental Microbiology, 1995, 63: 3789-3796.
[19] GOODFELLOW M, STACKERBRANDT E. Nucleic acid techniques in bacterial systematics [M]. New York: Wiley & Sons, 1991: 115-175.
[20] SAITOU N, NEI M. The neighbor-joining method: A new method for reconstructing phylogenetic trees [J]. Molecular Biology and Evolution, 1987, 4: 406-425.
[21] DE PEER V, Y, DE WACHTER R. TREECON for Windows: A software package for the construction and drawing of evolutionary trees for the Microsoft Windows environment [J]. Computer Applications in the Biosciences, 1994, 10: 569-570.
[22] MOYER C L, DOBBS F C, KARL D M. Estimation of diversity and community structure through restriction fragment length polymorphism distribution analysis of bacterial 16S rRNA genes from a microbial mat at an active, hydrothermal vent system, Loihi Seamount, Hawaii [J]. Applied and Environmental Microbiology, 1994, 60: 871-879.
Foundation item: Project(200805032) supported by Grands from Scientific Research Program of Marine Public Welfare Industry of China; Project (DYXM-115-02-2-07) supported by China Ocean Mineral Resources R&D Association (COMRA)
Received date: 2010-06-12; Accepted date: 2010-10-17
Corresponding author: ZHOU Hong-bo, Professor, PhD; Tel: +86-731-88877216; E-mail: zhouhb@mail.csu.edu.cn

