High level arsenic resistance in bacteria present in biooxidation tanks used to treat gold-bearing arsenopyrite concentrates: A review
来源期刊:中国有色金属学报(英文版)2008年第6期
论文作者:D. E. RAWLINGS
文章页码:1311 - 1318
Key words:Acidithiobacillus caldus; Leptospirillum; arsenic resistance genes; transposons; microbial consortia; biooxidation tanks
Abstract: The microbial consortium used in continuous-flow, stirred tank processes to treat gold-bearing arsenopyrite concentrates became adapted to high concentrations of arsenic over a number of years. The dominant microorganisms, Acidithiobacillus caldus and Leptospirillum ferriphilum, were found to contain two sets of arsenic resistance genes. One set of ars genes was present in all isolates of a species irrespective of whether they were highly arsenic resistant or not. A second set of ars genes was present on Tn21-like transposons and was found in all strains tested that had been adapted to high concentrations of arsenic. The arsenic resistance transposons present in At. caldus and L. ferriphilum were closely related, but sufficiently different for them to have been acquired independently rather than having been passed from one bacterium to the other. The transposons were transpositionally active in Escherchia coli and were shown to confer higher levels of arsenic resistance than the chromosomally-located ars genes where it was possible to test this. Transposons containing arsenic resistance genes that were identical or closely related to the transposon from L. ferriphilum, originally found in South Africa, were also found in both L. ferrooxidans and L. ferriphilum isolates from South America and Europe. An arsB gene knockout of At. caldus was produced by homologous recombination that demonstrated both the ability of the chromosomal ars genes to confer low levels of arsenic resistance in At. caldus and the development of a genetic system for the creation of knock-out mutants.

D. E. RAWLINGS
Department of Microbiology, University of Stellenbosch, Private Bag X1, Matieland, 7602, South Africa
Received 20 September 2008; accepted 5 November 2008
Abstract: The microbial consortium used in continuous-flow, stirred tank processes to treat gold-bearing arsenopyrite concentrates became adapted to high concentrations of arsenic over a number of years. The dominant microorganisms, Acidithiobacillus caldus and Leptospirillum ferriphilum, were found to contain two sets of arsenic resistance genes. One set of ars genes was present in all isolates of a species irrespective of whether they were highly arsenic resistant or not. A second set of ars genes was present on Tn21-like transposons and was found in all strains tested that had been adapted to high concentrations of arsenic. The arsenic resistance transposons present in At. caldus and L. ferriphilum were closely related, but sufficiently different for them to have been acquired independently rather than having been passed from one bacterium to the other. The transposons were transpositionally active in Escherchia coli and were shown to confer higher levels of arsenic resistance than the chromosomally-located ars genes where it was possible to test this. Transposons containing arsenic resistance genes that were identical or closely related to the transposon from L. ferriphilum, originally found in South Africa, were also found in both L. ferrooxidans and L. ferriphilum isolates from South America and Europe. An arsB gene knockout of At. caldus was produced by homologous recombination that demonstrated both the ability of the chromosomal ars genes to confer low levels of arsenic resistance in At. caldus and the development of a genetic system for the creation of knock-out mutants.
Key words: Acidithiobacillus caldus; Leptospirillum; arsenic resistance genes; transposons; microbial consortia; biooxidation tanks
1 Introduction
Continuous flow, stirred-tank based biooxidation processes for the recovery of gold from gold-bearing arsenopyrite concentrates were developed during the 1980s. A number of large-scale industrial plants that use these biooxidation process as a pre-treatment step prior to the recovery of gold by cyanidation have been built in several countries[1-2]. When arsenopyrite concentrates are biooxidized, levels of arsenic frequently reach saturation and soluble arsenate concentrations of 12 g/L and arsenite of 3-6 g/L have been reported[3]. During the early laboratory stages of development of the biooxidation process, oxidation of the arsenopyrite concentrate was slow and inefficient, partly because the bacteria were sensitive to the high concentrations of arsenic that were released. Arsenic inhibition was so severe that after a period of aeration, the mineral suspension was transferred to an unaerated tank where the pH was adjusted to 3.5 and the precipitated arsenic was removed by settling. The mineral suspension was then transferred to a fresh tank where aeration was continued. Initially more than one arsenic precipitation step was required. As the separation of arsenic from the insufficiently oxidized arsenopyrite was complicated, the process was uneconomic. During the operation of a continuous-flow tank aeration system, fast-growing arsenopyrite-biooxidizing cells displaced the more readily washed out slow-growing cells and this resulted in the selection of highly arsenic resistant microbes[4-5]. Over a period of about two years the bacterial consortium became sufficiently resistant to arsenic so that no arsenic precipitation steps were required. This enabled the residence time of the mineral concentrate in the series of aeration tanks to be reduced from about 12 to less than 7 days.
In 1986, the first commercial arsenopyrite biooxidation plant was built at the Fairview gold mine near the town of Barberton in South Africa. After three years of operation, not only had the total residence time of the mineral in the biooxidation tanks been reduced to a little over 3 days but also the quantity of mineral concentrate in suspension had been increased from 10% to 19% (w/v)[4]. Subsequently, several additional arsenopyrite biooxidation plants were built in other countries e.g. Brazil (Sao Bento), Australia (Wiluna and Harbour lights) and Ghana (Ashanti Sansu) Tamboraque (Lima, Peru) was built[1-2].
Continuous-flow tanks that are used for the biooxidation of arsenopyrite concentrates that operate at 40℃ are dominated by a mixture of the sulphur- oxidizing bacterium, Acidithiobacillus caldus, and the iron-oxidizing bacterium, Leptospirillum ferriphilum [6-7]. We wished to investigate what genetic changes had taken place that permitted a consortium of bacteria to increase their tolerance to total arsenic in solution from less than 1 g/L to 13 g/L. Unfortunately, no samples of the early biooxidation bacterial consortium were maintained and therefore it was not possible to compare the bacteria in the original arsenic-sensitive with the arsenic-tolerant consortia. However, isolates of the same species of bacteria as found in the biooxidation tanks that are unlikely to have been exposed to high levels of arsenic can be compared to the highly arsenic tolerant bacteria. In this work, investigations into how the At. caldus and L. ferriphilum strains present in arsenopyrite biooxidation tanks became highly arsenic resistant are reviewed.
2 Arsenic resistance mechanisms
In biological systems, arsenic typically exists in two oxidation states, arsenite (As Ⅲ) and arsenate (As Ⅴ). Arsenic resistance mechanisms have been reviewed extensively[8-11]. Typically, the minimal set of genes that confers resistance to both arsenite and arsenate are known as the arsR, arsB and arsC genes. The gene product, ArsR, is an arsenic-responsive gene regulator, ArsB, a membrane-located arsenite efflux pump and ArsC an arsenate reductase required to reduce arsenate to arsenite prior to its export from the cell by ArsB. Other frequently encountered arsenic resistance-associated genes are arsA and arsD. ArsA is an ATPase that forms a complex with ArsB and assists in the export of arsenite, while ArsD was reported to be a second regulator that controls the upper level of ars gene expression. More recently, ArsD has been found to act as an arsenite binding protein or metallochaperone[12] that sequesters intracellular arsenite and delivers it to ArsA before being pumped out of the cell through the action of the ArsAB membrane efflux pump (Fig.1).
3 Transposons containing arsenic resistance genes from At. caldus and L. ferriphilum
Strains of At. caldus and L. ferriphilum were isolated from the arsenopyrite biooxidation tanks at the Fairview mine and gene banks were constructed[3, 13]. The gene banks were transformed into an arsenic sensitive Escherichia coli strain ACSH50Iq and the transformants screened for arsenate and arsenite resistance[3, 13]. A Tn21-like transposon containing gene for arsenic resistance was isolated from both At. caldus and L. ferriphilum (Fig.1). The transposons were sequenced and named TnAtcArs and TnLfArs, respectively.
Fig.1 Comparison between genetic maps of transposons TnAtcArs from At. caldus and TnLfArs from L. ferriphilum (Numbers between transposons indicating percentage amino acid identity between equivalent gene products) (a), and diagrammatic representation of function of ars gene products (b)
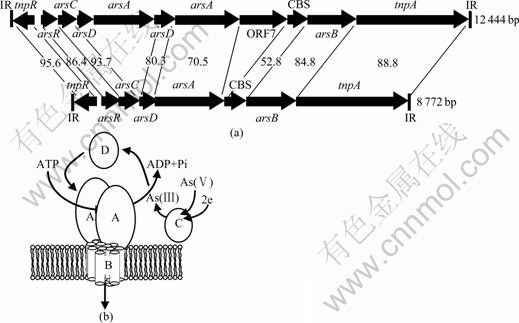
The transposon-associated arsenic resistance genes from At. caldus were contained on a 12 444 bp DNA fragment that has 40 bp long, Tn21-like inverted repeat(IR) sequences and genes for a Tn21 family-like resolvase (tnpR) and transposase (tnpA) (Fig.1). Seven ORFs closely related to arsenic resistance genes were located between the transposon resolvase and transposase genes. Arsenic resistance genes encoding products with high amino acid sequence homology to arsR, arsC, duplicate copies of arsD and arsA as well as an arsB were found. Between the second copy of arsA and the single copy of arsB, two open reading frames called ORF7 and ORF8 that had not previously been associated with ars genes were identified. The 470 aa long predicted product of ORF7 was reported to have the clear homology to the 466 aa NADH oxidase (46% identity/60% similarity) from the bacterium Lactobacillus plantarum. The product of ORF8 (158 aa) was reported to have the highest homology (54% identity/72% similarity) over approximately its entire length to what is known as the 146 aa CBS (cystathione- β-synthase)[14] domain of a membrane protein from Desulfitobacterium hafniense. Inactivation of either ORF7 or ORF8 did not appear to have a detectable effect on arsenite or arsenate resistance in E. coli and their function on TnAtcArs is not known.
The arsenic gene containing transposon, TnLfArs, isolated from L. ferriphilum was clearly related to TnAtcArs from At. caldus (Fig.1). The ars genes of TnLfArs were located between divergently transcribed tnpR and tnpA genes in a manner similar to the TnAtcArs transposon[13]. As for TnAtcArs, the ars genes of TnLfArs consisted of arsR, arsC, arsD, arsA and arsB all transcribed in the same direction. Between arsA and arsB there was an open reading frame equivalent to ORF8 of TnAtcArs. Despite the overall similarities, there were some important differences between TnLfArs and TnAtcArs. TnLfArs was only 8 772 bp in length compared with 12 444 bp for TnAtcArs. The main reason for the size difference was that ORF7 and the arsDA duplication of TnAtcArs were absent from TnLfArs. The TnLfArs 43 bp left and right invert repeats (IR) differed from one another by 1 bp and were 3 bp longer than those for TnAtcArs. Furthermore, although the amino acid sequences of several gene products were highly conserved (TnpR, ArsC and TnpA had 96%, 94% and 89% amino acid sequences with identity, respectively), others were much less conserved. For example, ORF8 and ArsA had only 53% and 71% amino acid sequence identity, respectively (Fig.1).
The differences between the two ars transposons suggested that it was highly unlikely that the transposons had been passed from At. caldus to L. ferriphilum or vice versa in the biooxidation tank environment. The differences in DNA sequences between the genes and the amino acid sequences of their products were much too substantial for these to have arisen in the ten to fifteen years or less since the biooxidation plant at Fairview mine began operation and the time of sample collection. This suggested that although the two types of bacteria had acquired related transposons to increase their arsenic resistance, they had acquired the ars transposons independently of each other.
4 TnAtcArs and TnLfArs were transpo- sitionally active but no evidence was obtained that transposition frequency was increased in presence of arsenic
Both TnAtsArs and TnLfArs were tested to see whether they were still transpositionally active[3, 13] in E. coli by using plasmid pSa as a ‘transposon trap’. A diagramatic representation of the experiments is shown in Fig.2. TnAtcArs was cloned into a non-conjugative plasmid (lacked an oriT origin of transfer) and it was shown that when present on its own in a cell, TnAtcArs could not be conjugated to an E. coli rifampicin resistant recipient. However, when the conjugative plasmid pSa (chloramphenicol resistant) was present in the same cell as the non-mobilizable plasmid containing TnAtcArs, the TnAtcArs transposon could be conjugated into an E. coli rifampicin resistant mutant (Fig.2). The explanation for this was that TnAtcArs had transposed into the pSa ‘transposon trap’ in the donor strain and that this pSa-TnAtcArs plasmid had been transferred by conjugation to the E. coli recipient. Transposition of TnAtcArs into pSa was confirmed by digestion of pSa and pSa-TnAtcArs with the restriction endonuclease enzymes BglⅡ and HindⅢ. It was found that size of one of the DNA fragments of pSa-TnAtcArs was larger than pSa. Furthermore, whereas pSa gave no hybridization signal when using arsB of TnAtcArs as probe, the pSa-TnAtcArs plasmid did give a positive hybridization signal, indicating that the increase in size was due to the acquisition of arsenic resistance genes.
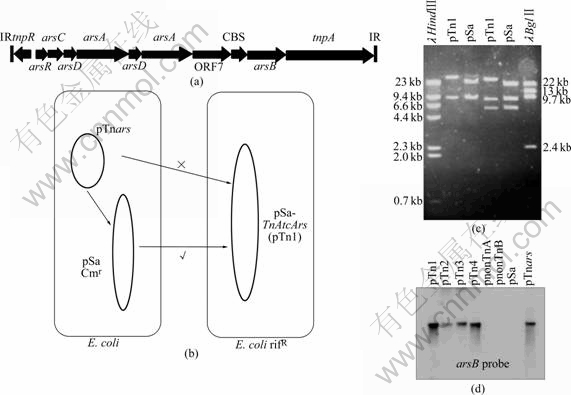
Fig.2 Representation of TnAtcArs inserted into plasmid pUC19 to produce non-mobilizable construct, pTnars (a); Diagrammatic illustration of experiment of TnAtcArs present on pTnars transposed into conjugative plasmid, pSa, before transfer to E. coli rifampicin resistant recipient (b); Restriction endonulease analysis of pSa before and after insertion of TnAtcArs (pTn1) (c); Southern hybridization of pTn1 and three similar transposon insertions into pSa (pTn2-4) probed with At. caldus arsB (Lanes marked pnonTnA and pnonTnB containing plamids from transconjugants where donor cells contained pSa but not pTnars) (d)
Similar results were obtained when TnLfArs was tested for transposition activity in E. coli using the same type of experiment, indicating that both TnAtsArs and TnLfArs were transpositionally active[13].
Both transposons TnAtsArs and TnLfArs appeared to be dedicated to the transposition of the arsenic resistance genes located between the tnpR (resolvase) and tnpA (transposase) genes. Furthermore, the gene for the transposase was located downstream of the arsB gene and was transcribed in the same direction as the ars genes. This raised the question of whether transcriptional read-through from the ars genes might increase the level of expression of the tnpA gene and hence the frequency of transposition might increase in response to arsenic. An increased frequency of transposition in response to mercury has been reported in the mercury resistance transposon Tn501[15]. However, we were unable to detect an increase in the frequency of transposition in response to arsenic in E. coli. This was possibly due to a fairly large scatter in the transposition frequency but northern hybridisation data showed that tnpA expression was not affected by the presence of arsenite, and therefore, increased transposition in response to arsenite probably does not occur[3].
5 Not all strains of At. caldus and L. ferriphilum contain ars resistance transposon
We had obtained six strains of At. caldus, three of which had been previously exposed to arsenic as they had previously been isolated from arsenopyrite concentrate biooxidation tanks. Although one strain was from a nickel ore biooxidation tank, the inoculum had originated from an arsenopyrite biooxidation tank some years previously. In addition, we obtained three strains that were unlikely to have been exposed to high levels of arsenic. These originated from Birch Coppice (BC13) and a coal spoil (KU) in the United Kingdom and from a bioreactor in Brisbane, Australia (C-SH12). All strains of At. caldus previously exposed to arsenic were found to contain the TnAtcArs operon whereas none of the three strains not known to have been previously exposed to arsenic had the arsenic resistance transposon[16-17]. However, all six At. caldus strains contained chromosomally-encoded arsenic resistance genes.
A similar situation appeared to exist in Leptospirillum. Genomic DNA from three L. ferrooxidans strains (ATCC49879, DSM2705 and Chil-Lf2) and three L. ferriphilum strains (ATCC49881, Fairview and Warwick) was analysed for the presence of the transposon and chromosomal ars genes. When a probe with a fragment specific for TnLfArs was used in Southern hybridisation experiments, only L. ferriphilum Fairview gave a signal[13]. Both L. ferriphilum and L. ferrooxidans have chromosomal-type arsenic genes in addition to the Tn-related genes [18]. The L. ferriphilum chromosomal ars genes are atypical in that the arsB gene is preceded by an arsR and arsC and these two genes appeared to have been fused[13]. When the arsRCB chromosomal genes of L. ferriphilum was used as probe, all of the L. ferriphilum strains gave a hybridisation signal, but none of the L. ferrooxidans strains, even under lower stringency conditions. This suggested that the chromosomal ars genes of L. ferriphilum and L. ferrooxidans had dissimilar DNA sequences. In subsequent experiments, the chromosomal arsB gene from L. ferrooxidans was isolated[18]. In L. ferrooxidans, the arsB appears to occur as a single gene, with a predicted amino acid sequence that is substantially different from L. ferriphilum. The absence of a gene for an ArsR regulator linked to arsB raised the question of how arsB expression is regulated in L. ferrooxidans. The finding suggested that either a gene for an arsR regulator lies elsewhere on the L. ferrooxidans chromosome or that the arsB gene that was isolated may be redundant.
6 Transposon encoded arsenic resistance genes confer much higher levels of arsenic resistance than chromosomally encoded arsenic resistance genes
The transposon-encoded arsenic resistance genes appeared to confer much higher levels of arsenic resistance than the chromosomally encoded arsenic resistance genes. Since no strains of At. caldus lacking the chromosomal arsenic resistance genes existed at the time, the levels of arsenic resistance were tested in E. coli after cloning the two types of resistance genes in an E. coli strain that lacked the usual chromosomal arsenic resistance genes[3, 19]. Whereas TnAtcArs conferred resistance to E. coli ars mutants of at least 5 mmol/L arsenite, the cloned chromosomal resistance genes conferred resistance to only 0.25 mmol/L arsenite. Resistance to arsenate was more complicated to compare because of interference by phosphate in the growth medium.
That an increase in arsenic resistance was conferred by TnAtcArs was also tested in the normal host. At. caldus strain C-SH12 that lacked TnAtcArs grew in medium containing up to approximately 20 mmol/L arsenite (Fig.3). After the transfer of TnAtcArs to strain C-SH12 by conjugation, the transconjugant strain containing both the chromosomal- and transposon- encoded ars genes, grew in thiosulfate medium containing greater than 50 mmol/L arsenite at which point it was difficult to maintain arsenite in solution[19]. This illustrated the effectiveness of transposon-located ars genes in conferring arsenic resistance relative to a strain containing only the chromosomal ars genes.
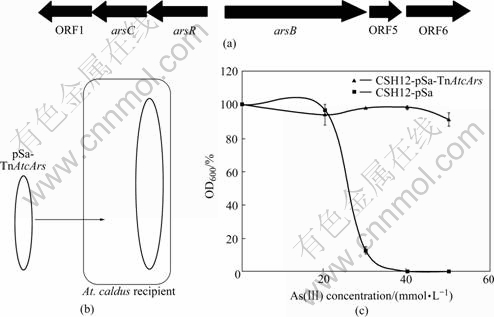
Fig.3 Genetic map of region including chromosomal ars genes of At. Caldus (a); Plasmid pSa-TnAtcArs transferred by conjugation from E. coli to At. caldus (b); Growth after 19 d of inclubation of At. caldus strain C-SH12 containing only pSa compared with growth of At. caldus strain C-SH12 containing pSa-TnAtcArs in presence of increasing concentrations of arsenite (c) [19]
With respect to L. ferriphilum, although the genes on TnLfArs conferred arsenic resistance to E. coli when transferred to that bacterium, the cloned chromosomal genes were not functional in E. coli[13]. The relative effectiveness of the transposon-located and chromosome- located ars genes could therefore not be tested. Nevertheless, growth in arsenic of the L. ferriphilum Fairview strain containing TnLfArs, was compared with growth in arsenic of the L. ferriphilum type strain ATCC49881 which lacked TnLfArs. L. ferriphilum ATCC49881 grew in iron medium containing arsenic up to approximately 40 mmol/L of either As(Ⅴ) or As(Ⅲ) [13]. In contrast, L. ferriphilum Fairview was capable of growth and iron oxidation of up to 60 mmol/L of either As(Ⅴ) or As(Ⅲ). At higher concentrations it was difficult to keep the arsenic in solution and the resistance of this strain may therefore be higher. Since two different L. ferriphilum strains were used, this is not proof that the TnLfArs conferred increased arsenic resistance, but suggests that this might be the case.
7 Distribution of TnLfArs genes
As described above, it was found that the strains of At. caldus and L. ferriphilum that dominate the gold-bearing arsenopyrite biooxidation tanks at the Fairview mine had acquired related but different transposons of the Tn21-family that contained an operon that conferred arsenic resistance. Furthermore, only the three At. caldus strains and the L. ferriphilum Fairview strain that had previously been exposed to high levels of arsenic, contained an arsenic resistance transposon[13].
Many of the commercial arsenopyrite biooxidation processes that have been built in different parts of the world were inoculated with the microbial consortium from the biooxidation tanks of the Fairview mine. An exception to this is the biooxidation plant at Tamboraque (near Lima, Peru) where the consortium used in those tanks originated from a mixture of locally available iron and sulfur oxidizing microorganisms[20]. We wished to determine whether arsenic resistance in this consortium had occurred through the acquisition of a similar transposon as the Fairview consortium or by a different mechanism. Unfortunately we were not able to isolate a viable culture of a sulfur-oxidizing microorganism from the Tamboraque mineral sample following its transport to South Africa from Peru. However, we were able to isolate an iron oxidizer that turned out to be a strain of L. ferrooxidans rather than a L. ferriphilum[18]. We also had access to several isolates of both L. ferriphilum and L. ferrooxidans from Europe and these were also screened to detect whether they contained an arsenic resistance transposon related to those present in the Fairview strain even though they had not been selected for enhanced arsenic resistance. PCR and Southern hybridization experiments indicated that arsenic resistance genes that were identical or closely related to those from South Africa were found in both L. ferrooxidans and L. ferriphilum isolates from South America and Europe [18].
The widespread occurrence of arsenic resistance transposons amongst different isolates of L. ferriphilum and L. ferrooxidans from at least three continents (Africa, South America and Europe) suggested that it should be possible to select for highly arsenic resistant biomining microorganisms from many different sources and that it was therefore probably unnecessary to import pre-adapted, arsenic resistant consortia.
8 Origin of Tn21-like arsenic resistance transposons
Since the TnAtcArs and TnLfArs transposons were transpositionally active and not present in all strains of At. caldus, L. ferriphilum or L. ferrooxidans, these transposons would be expected to have been acquired from the horizontal gene pool. This raised the question of whether they originated in bacteria such as the At. caldus and Leptospirillum isolates described here, or whether the arsenic resistance transposons originated in other organisms and were acquired by the ‘biomining’ bacteria by horizontal gene transfer.
A BLAST search using the L. ferriphilum ars genes indicated that they were most similar to transposon- related ars genes identified in Methylobacillus flagellatus [21] and Alcaligenes faecalis (AY297781)[13]. Even though this work was reported in 2006, the ars genes from the transposons found in M. flagellatus and A. faecalis remain the most closely related ars genes in sequence databases. A comparison of the structure of the TnLfArs ars transposon with that of A. faecalis and Methylobacillus flagellatus is shown in Fig.4 (structures of TnAtcArs and TnLfArs were compared in Fig.1). In general, the ars operon structures of TnLfArs, TnAtcArs and A. faecalis transposons are closely related. Exceptions are that TnAtcArs has an additional NADH oxidoreductase-like gene (ORF7) and in addition, the M. flagellatus arsR and arsC genes appear to have been fused. A comparison of the sequence identity of the ars gene products of the M. flagellatus and A. faecalis transposons with those of TnLfArs is also made in Fig.4. Some of predicted protein products within the ars operons are highly related to each other. For example, the ArsD, ArsA and ArsB proteins have 97.5%, 94.6% and 97.4% amino acid sequence identity, respectively.

Fig.4 Diagrammatic representation of TnLfArs compared to related transposons found in Alcaligenes faecalis (AY297781) and Methylobacillus flagellatus (Numbers between transposons indicating percentage amino acid identity between equivalent ars gene products) [13]
With respect to the rest of the transposon structures, the A. faecalis transposon appears to have undergone a deletion of tnpR and an inverted repeat. The structure of the M. flagellatus transposon is different in that the tnpA and tnpR genes are adjacent to each other as for most other Tn21-subfamily (and Tn3-family) transposons where they form a functional unit[22].
Together these results suggest that there exists a group of related Tn21-like transposons that carry arsenic resistance genes member of which are present in a number of microorganisms, most of which have still to be discovered[13]. Given current knowledge it is not yet possible to determine from which organisms TnAtcArs and TnLfArs originated or whether either transposon is the source of the related group of transposons. Futher discoveries of transposons from this group might clarify this.
9 Construction of At. caldus chromosomal ars gene knockout mutants
Arsenic resistance genes have proved useful in another context, of that a protocol for creating At. caldus chromosomal gene knockout mutants is established. The usefulness of the At. caldus chromosomal ars genes was that the creation of a knockout mutant was not expected to be lethal to the host. To create an arsB knockout mutant, the arsB gene of At. caldus was cloned into a At. caldus suicide cloning vector. This vector contained the origin of transfer (oriT) sequence of the broad-host-range conjugative plasmid, pSa, and could replicate in E. coli but not in At. caldus[23]. The cloned arsB gene was inactivated by the deletion of a small piece of the gene and replacing it with a kanamycin resistance gene. This construct containing the inactivated arsB gene was transferred by conjugation from a strain of E. coli containing the conjugative plasmid pSa to At. caldus. Following conjugation, the only means by which the marker on the suicide plasmid could be recovered was by homologous recombination with the ars genes present on the At. cadlus chromosome. Transconjugants were selected on a thiosulfate medium containing kanamycin and those grown were screened using Southern hybridization with a probe made from the kanamycin resistance gene. This confirmed that the kanamycin gene had integrated into the At. caldus chromosome probably by means of the inactivated arsB chromosomal gene. As both double and single cross-over insertions within the arsB gene were possible, the presence of single cross- over mutants was screened by Southern hybridization using the small fragment deleted from the arsB gene as probe[23]. This deleted fragment would be missing from double cross-over but not single cross-over mutants. Of the 140 kanamycin resistant colonies checked, only six were double cross-over mutants where the wild type arsB gene had been replaced by the inactivated arsB gene. The other At. caldus mutants were single cross-over mutants where the conjugated suicide plasmid had been integrated into the chromosome such that the transconjugants contained both an active and an inactive copy of the arsB gene. When tested for arsenic resistance, isolates containing a single inactivated arsB gene were extremely sensitive to arsenite. This indicated that arsB gene knockouts had been successfully made and that the chromosomal At. caldus ars genes contributed to low-level arsenic resistance[23].
10 Conclusions
1) All strains of At. caldus, L. ferriphilum and L. ferrooxidans tested contain ars genes present on their chromosomes. In the case of At. caldus, these were shown to be functional in both At. caldus and E. coli. The chromosomal genes of both species of Leptospirillum were atypical and non-functional in E. coli. It is uncertain whether these chromosomal genes are functional in Leptospirillum. This is especially so in the case of L. ferrooxidans where only a single arsB was found without any sign of an arsC gene (arsenate reductase) or arsR regulator.
2) Beside chromosomal ars genes, At. caldus, L. ferriphilum and L. ferrooxidans strains isolated from biooxidation tanks treating gold-bearing arsenopyrite ores contain a second set of arsenic resistance genes that are present on the Tn21-like transposons, TnAtcArs and TnLfArs. In the case of At. caldus, these genes conferred high levels of resistance to arsenite and arsenate in both At. caldus and E. coli. In the case of L. ferriphilum, high levels of arsenic resistance occurred when the TnLfArs transposon was present in E. coli, but gene transfer techniques did not exist to test to what extent arsenic resistance was enhanced in strains of Leptospirillum that lacked the transposon.
3) Both the TnAtcArs and TnLfArs transposons are transpositionally active in E. coli, but no evidence was obtained that the rate of transposition was increased in the presence of arsenic.
4) The TnLfArs transposon or a transposon highly similar to TnLfArs is present in isolates of L. ferriphilum and L. ferrooxidans from at least three continents including some isolates not known to have been previously exposed to high levels of arsenic. This suggests that it should be possible to isolate consortia of arsenic resistant microorganisms without having to resort to obtaining such consortia from plants that have previously been exposed to arsenic such as commercially operating arsenopyrite biooxidation plants.
5) Although it is not yet possible to determine from where the Tn21-like ars gene containing transposons originated, a family of related transposons appears to exist in a variety of bacteria including typical heterotrophic microorganisms.
6) At. caldus strains in which the chromosomal ars genes were inactivated using a kanamycin resistance gene became highly sensitive to arsenic. This demonstrated that the chromosomal ars genes conferred not only the low levels of arsenic resistance to At. caldus, but also the development of a genetic system for the creation of knock-out mutants in At. caldus.
Acknowledgements
The author thanks the University of Stellenbosch, the National Research Foundation (Pretoria) and the BioMinE project 500329 of the EU framework 6 for sponsoring this work.
References
[1] DEW D W, LAWSON E N, BROADHURST J L. The BIOX? process for biooxidation of gold-bearing ores or concentrates [C]// RAWLINGS D E, ed. Biomining: Theory, Microbes and Industrial Processes. Berlin: Springer-Verlag, 1997: 45-80.
[3] TUFFIN I M, DE GROOT P, DEANE S M, RAWLINGS D E. An unusual Tn21-like transposon containing an ars operon is present in highly arsenic resistant strains of the biomining bacterium Acidithiobacillus caldus [J]. Microbiology, 2005, 151: 3027-3039.
[4] RAWLINGS D E, SILVER S. Mining with microbes [J]. Bio/Technology, 1995, 13: 773-779.
[5] RAWLINGS D E, JOHNSON D B. The microbiology of biomining: Development and optimization of mineral-oxidizing microbial consortia [J]. Microbiology, 2007, 153: 315-324.
[6] RAWLINGS D E, CORAM N J, GARDNER M N, DEANE S M. Thiobacillus caldus and Leptospirillum ferrooxidans are widely distributed in continuous flow biooxidation tanks used to treat a variety of ores and concentrates [C]// AMILS R, BALLESTER A, eds. Biohydrometallurgy and the Environment Toward the Mining of the 21st Century, Part A. Amsterdam: Elsevier, 1999: 777-786.
[7] CORAM N J, RAWLINGS D E. Molecular relationship between two groups of the genus Leptospirillum and the finding that L. ferriphilum sp. nov. dominates South African commercial biooxidation tanks that operate at 40℃ [J]. Applied and Environmental Microbiology, 2002, 68: 838-845.
[8] CERVANTES C, JI G, RAM?REZ J L, SILVER S. Resistance to arsenic compounds in microorganisms [J]. FEMS Microbiological Reviews, 1994, 15: 355-367.
[9] SILVER S, PHUNG L T. Bacterial heavy metal resistance: New surprises [J]. Annual Review of Microbiology, 1996, 50: 753-789.
[10] ROSEN B P. Families of arsenic transporters [J]. Trends in Microbiology, 1999, 7: 207-212.
[11] MUKHOPADHYAY R, ROSEN B P, PHUNG L T, SILVER S. Microbial arsenic: From geocycles to genes and enzymes [J]. FEMS Microbiological Reviews, 2002, 26: 311-325.
[12] LIN Y, WALMSLEY A R, ROSEN B P. An arsenic metallochaperone for an arsenic detoxification pump [J]. Proceedings of the National Academy of Science (USA), 2006, 103: 15617-15622.
[13] TUFFIN I M, HECTOR S B, DEANE S M, RAWLINGS D E. The resistance determinants of a highly arsenic resistant strain of Leptospirillum ferriphilum isolated from a commercial biooxidation tank [J]. Applied and Environmental Microbiology, 2006, 72: 2247-2253.
[14] BATEMAN A. The structure of a domain common to archaebacteria and the homocystinuria disease protein [J]. Trends in Biochemistry, 1997, 22: 12-13.
[15] SCHMITT R, ALTENBUCHNER J, GRINSTED J. Complementation of transposition functions encoded by transposons Tn501 (Hg) and Tn1721 (Tc) [C]// LEVY S B, CLOWES R C, KOENIG E C, eds. Molecular Biology, Pathogenicity and Ecology of Bacterial Plasmids. New York: Plenum, 1981: 359-370.
[17] TUFFIN M, DE GROOT P, DEANE S M, RAWLINGS D E. Multiple sets of arsenic resistance genes are present within highly arsenic resistant industrial strains of the biomining bacterium, Acidithiobacillus caldus [J]. International Congress Series, 2004, 1275: 165-172.
[19] KOTZE A A, TUFFIN I M, DEANE S M, RAWLINGS D E. Cloning and characterisation of the chromosomal arsenic resistance genes from Acidithiobacillus caldus and enhanced arsenic resistance on conjugal transfer of ars genes located on transposon TnAtcArs [J]. Microbiology, 2006, 152: 3551-3560.
[20] LOAYZA C, LY M E, YUPANQUI R, ROM?N G. Laboratory biooxidation tests of arsenopyrite concentrate for the Tamboraque industrial plant [C]// AMILS R, BALLESTER A, eds. Biohydrometallurgy and the Environment Toward the Mining of the 21st Century, Part A. Amsterdam: Elsevier, 1999: 405-410.
[21] CHISTOSERDOVA L, LAPIDUS A, HAN C, GOODWIN L, SAUNDERS E, BRETTIN T, TAPIA R, GILNA P, LUCAS S, RICHARDSON P M, LIDSTROM M E. Genome of Methylobacillus flagellatus, molecular basis for obligate methylotrophy, and polyphyletic origin of methylotrophy [J]. Journal of Bacteriology, 2007, 189: 4020-4027.
[22] LIEBERT C A, HALL R M, SUMMERS A O. Transposon Tn21, flagship of the floating genome [J]. Microbiology and Molecular Biology Reviews, 1999, 63: 507-522.
[23] VAN ZYL L J, VAN MUNSTER J M, RAWLINGS D E. Construction of arsB and tetH mutants of the sulfur-oxidizing bacterium, Acidithiobacillus caldus by marker exchange [J]. Applied and Environmental Microbiology, 2008, 74: 5686-5694.
Corresponding author: D. E. RAWLINGS; E-mail: der@sun.ac.za
(Edited by YANG Bing)

