
J. Cent. South Univ. (2020) 27: 1386-1394
DOI: https://doi.org/10.1007/s11771-020-4374-2

Comparison of bioleaching of chalcopyrite concentrates with mixed culture after cryopreservation with PEG-2000 in liquid nitrogen
WU Xue-ling(吴学玲)1, 2, WU Xiao-yan(吴晓燕)1, DENG Fan-fan(邓凡凡)1, SHEN Li(申丽)1, 2,
LI Jiao-kun(李交昆)1, 2, PENG Tang-jian(彭堂见)1, 2, YU Run-lan(余润兰)1, 2, LIU Yuan-dong(刘元东)1, 2,
CHEN Miao(陈淼)3, QIU Guan-zhou(邱冠周)1, 2, ZENG Wei-min(曾伟民)1, 2
1. School of Minerals Processing and Bioengineering, Central South University, Changsha 410083, China;
2. Key Laboratory of Biohydrometallurgy of Ministry of Education, Central South University,Changsha 410083, China;
3. CSIRO Process Science and Engineering Clayton, Victoria, 3169, Australia
 Central South University Press and Springer-Verlag GmbH Germany, part of Springer Nature 2020
Central South University Press and Springer-Verlag GmbH Germany, part of Springer Nature 2020
Abstract: A mixed culture of bioleaching microorganisms was enriched. Then the mixed culture was preserved by liquid nitrogen cryopreservation for 6 months and 12 months, respectively, using PEG-2000 as the protective agent. The chalcopyrite leaching ability, activity and diversity of the mixed culture before and after preservation were compared. The results showed that the copper extraction rate was 95.7% in chalcopyrite bioleaching within 20 d by the original culture. After cryopreservation for 6 months and 12 months, the copper extraction rate of the mixed culture was 94.9% within 25 d and 93.6% within 35 d, respectively. The cell viability achieved 87% and 41% after being preserved for 6 months and 12 months, respectively. Furthermore, the ecology analysis identified Acidithiobacillus ferrooxidans, Acidithiobacillus caldus, Sulfobacillus thermotolerans and Pseudomonas aeruginosa in the original mixed culture. After cryopreservation for 12 months, the composition of community changed, but the predominant microorganisms still existed.
Key words: mixed cultures; chalcopyrite bioleaching; PEG-2000; cryopreservation; microbial community
Cite this article as: WU Xue-ling, WU Xiao-yan, DENG Fan-fan, SHEN Li, LI Jiao-kun, PENG Tang-jian, YU Run-lan, LIU Yuan-dong, CHEN Miao, QIU Guan-zhou, ZENG Wei-min. Comparison of bioleaching of chalcopyrite concentrates with mixed culture after cryopreservation with PEG-2000 in liquid nitrogen [J]. Journal of Central South University, 2020, 27(5): 1386-1394. DOI: https://doi.org/10.1007/s11771-020-4374-2.
1 Introduction
The utilization of low grade ore, tailings and waste rock has drawn lots of attention with increasing demand for metal resources. Biohydrometallurgy technology was reported as an effective way to treat mineral resources due to its low cost, relatively low pollution and high adaptability [1, 2]. The microorganism is the key factor to improve the biohydrometallurgy process. The mixed culture compared with pure culture could play a more important role because mixed culture may have cooperation mechanism, leading to synergistic effect of bioleaching process [3-5]. For example, Acidithiobacillus caldus is a sulphur-oxidizing bacteria, and when it cooperated with Acidithiobacillus ferrooxidans, which is a sulphur- and ferrous ion-oxidizing bacterium, a good performance for chalcopyrite bioleaching was achieved [6]. RAMANATHAN et al [7] found that the bioleaching of Cu from municipal solid waste incineration fly ash using a mixed culture of the four isolates (Agromyces aurantiacus TRTYP3, Alkalibacterium pelagium TRTYP5, Alkalibacterium sp. TRTYP6 and Bacillus foraminis TRTYP17) was more effective than pure cultures or mixtures of two or three isolates. NIE et al [8] also used mixed culture of A. ferrooxidans, Pseudomonas sp., A. acidophilum, and Sphingomonas sp. to leach copper and nickel and their results showed that the bioleaching ratios of copper and nickel with mixed culture were 65% and 56%, respectively, which were higher than those with the single strain (41% and 36% only using A. ferrooxidans).
The strains in the mixed cultures undergo mutations during frequent passages and their bioleaching capacity is reduced [9]. Therefore, to preserve the high efficiency of mixed cultures becomes an important issue. In addition, an appropriate selection of protective agents is essential during the cryopreservation [10][Li, 2011 #128]. According to previous reports, polyethylene glycol can be used as universal cryoprotectant for different species of bacteria in long-time preservation [11]. Polyethylene glycol has been used in cryomicrobiology with concentration of 5%-45% and it is available with molecular weight ranging between 200 and 40000 [12]. For a combine use of PEG-6000 and 10% Me2SO, the cryopreservation efficiency of the algae P. yezoensis was very good [13]. In our previous study, it was found that PEG-2000 was a desirable cytoprotective agent to cryopreserve the pure bioleaching strain (Patent 201210559004.4 on the site http://www.soopat. com/). Furthermore, our previous study found that A. ferrooxidans has the highest survival rate and the shortest time to recover cell activity when 20% PEG-2000 was used as protectant agent [14]. We have also studied the effect of PEG-2000, PEG-12000, PEG-20000, and glycerol as the protective agent on the survival rate of Acidithiobacillus caldus during liquid nitrogen preservation in the earlier, and the results showed that it has the highest survival rate (90.71%) when using PEG-2000 as protective agent [9]. In recent years, lots of high efficient mixed cultures for bioleaching have been enriched [5, 8]. However, there are few reports about the cryopreservation of the mixed bioleaching culture. Furthermore, it is poorly understood about the microbial shift in the cryopreservation process and how such shift influences leaching capacity.
Therefore, in this study, PEG-2000 was used to cryopreserve the mixed culture with good performance of chalcopyrite bioleaching. The effects of cryopreservation on the bioleaching ability and the microbial community were investigated. These will be beneficial to find a good preservation way for the mixed bioleaching culture.
2 Materials and methods
2.1 Culture medium and microorganisms
A mixed culture of acidophilic microorganisms was collected from a bioleaching heap at Dexing Copper Mine in Jiangxi Province, China. The pH and temperature of the original sample were 2.5 and 32 °C, respectively. The AMD sample was filtered by 0.2 μm nylon filtration membranes (Biobasic Inc.), and then the mixed culture was collected and preserved at 4 °C. The mixed culture was cultured in the 9K medium with 30 g/L FeSO4・7H2O, 10 g/L S and 2% (m/V) chalcopyrite as the energy source. The initial pH was 2.0, temperature was 40°C, and shaking speed was 170 r/min. After passage culture for 6 d, the biomass reached 1×109 cell/mL, and microorganisms were collected by centrifugation and used for bioleaching, cryopreservation and ecology analysis experiments.
2.2 Minerals composition
The chalcopyrite concentrate was obtained from Meizhou in Guangdong Province, China. X-ray diffraction (XRD) analysis showed that chalcopyrite (60%) and pyrrhotite (25%) were the major components and galena (5%), quartz (3%), and sphalerite (2%) were the minor ones. The main chemical composition of the concentrate was 19% Cu, 24% Fe, 37% S, 3% Pb, and 5% Zn. The mineral was ground till the particle size <75 μm.
2.3 Bioleaching experiments
Bioleaching experiments were carried out in 250-mL shake flasks containing 100 mL 9K medium with 2% pulp concentration at 30 °C, 170 r/min, and initial pH 2.0. The mixed culture (enriched in Section 2.1) was inoculated into shake flasks. The initial cell density was adjusted to 1×107 cell/mL. Distilled water was added into the flasks every day in order to compensate for evaporation losses. The concentrations of Cu2+, Fe2+, and Fe3+ were analyzed by atomic absorption spectrophotometry. pH value was measured by precision acidity meter (pHS-3C). Cell density was analyzed by using blood counting chamber under optical microscope. All experiments were repeated in triplicates, using aseptic medium without inoculation as a blank control.
2.4 Cell preservation, recovery and bioleaching
The microorganisms collected in Section 2.1 were concentrated to a density of 2×1010 cell/mL for liquid nitrogen cryopreservation. PEG-2000 was added into the cell suspension as a cryoprotectant at a concentration of 20% (W/V). In the comparative experiments, 30% glycerol was added as the cryoprotectant. All cell samples were pre-frozen in a refrigerator at -20 °C for 2 h, and then transferred to a refrigerator at -70 °C. After 2 h, they were taken out and quickly transferred to a liquid nitrogen tank for cryopreservation [15, 16].
When microorganisms were preserved for 6 and 12 months, the frozen mixed culture was revived by thawing for 10 min at 37 °C in a water bath, and then transferred to a sterile Eppendorf. The preserved culture was then centrifuged at 10000g for 3 min, and then the supernatant was removed. The sediment was washed with sterile water three times to remove the protective agent, and inoculated in 9K medium for 6 d, using 30 g/L FeSO4・7H2O and 10 g/L S as the energy source. During cultivation, the cell density was analyzed by using blood counting chamber under optical microscope and the cell viability was measured according to ZENG et al [17]. Cell viability rate (v) was measured by cell density counted before preservation (N0) and after different times of storage (Nt). The viability rate (v) was expressed by the cell survival rate that was calculated: v=100%×Nt/N0, where N0 is the cell density for the original culture growing at the 6th day, and Nt is the cell density for the preserved culture growing at the 6th day. Meanwhile, the cells growing at the 6th day were collected for RFLP analysis.
2.5 RFLP analysis from both original and preserved cultures
2.5.1 DNA extraction and purification
Genomic DNAs from original culture and preserved cultures after 6 and 12 months were extracted using bacteria genomic DNA extraction kit (Tiangen Biotech Co., Ltd., Beijing, China). The 16S rDNA sequences were amplified with the bacteria universal primers 27F and 1492R as described by FU et al [18]. The resulting PCR products were purified with a commercial kit (Gel Extraction Kit, Tiangen Biotech Co. Ltd., Beijing, China).
2.5.2 PCR amplification and cloning of 16S rDNA sequences
The purified amplicons of 16S rDNA sequences were cloned into the pGM-T vector and transferred into Escherichia coli Top10F competent cells according to the manufacturer’s instructions (Tiangen Biotech Co. Ltd., Beijing, China). Positive clones were identified based on blue-white screening. White colonies from three libraries were randomly selected and the cloned inserts were re-amplified by PCR with the vector-specific T7 (5’-ATT TAG GTG ACA CTA TAG-3’) and SP6 (5’-TAA TAC GAC TCA CTA TAG GG-3’) primers, with an initial denaturation at 94 °C for 5 min, followed by 32 cycles at 94 °C for 45 s, 52 °C for 45 s, and 72 °C for 1 min, and completed with an extension period of 10 min at 72 °C. Next, the re-amplified products from each library were digested by using the restriction endonuclease Hin61 and MspI (Fermentas, MBI) at 37 °C overnight. The system for the reaction was described by FU et al [18]. The digestion products were separated by gel electrophoresis (3 h, 80 V) in 3.0% low-melting-point agarose in tris acetate buffer and analyzed by staining with EB under UV illumination. The banding patterns were grouped into an operational taxonomic unit (OTU) based on the DNA banding pattern of individual clones. Each banding pattern constituting one distinct OTU was based on how the restriction enzyme cuts the PCR product and the representative clones were selected for sequencing.
Sequencing and phylogenetic analysis unique OTUs were sequenced by the GenScript (Nanjing) Co., Ltd., and the sequences were initially estimated using the BLAST at the National Center for Biotechnology Information (http://www.ncbi.nlm. nih.gov/BLAST/). The closest 16S rDNA sequences were aligned with CLUSTALX 1.83. A phylogenetic tree was constructed by the neighbor joining method, using MEGA 5.05, and the robustness of the phylogeny was tested by a bootstrap analysis with 1000 iterations.
2.6 Statistical analysis
The experimental results were expressed as the means ± standard error of the three replicates. One-way analysis of variance (ANOVA) was carried out to confirm the variability of data and validity of results, and differences were considered significant at p<0.05 (SPSS 19.0 for Windows).
3 Results and discussion
3.1 Liquid nitrogen preservation of mixed culture using PEG-2000 as cryoprotectant
Liquid nitrogen cryopreservation has been used to preserve microorganisms for decades and it is one of the preferred methods. The protective agent is one of the key factors to determine the cell viability of preserve culture. In this work, PEG-2000 was indicated as an effective cryoprotectant to preserve the mixed bioleaching culture, and especially to protect the activity of the predominant bioleaching microorganisms. This is because PEG-2000 is a kind of permeating agents [19], and it can permeate into the cell-wall and form a buffer layer between the cell wall and membrane. This layer could avoid inner cell to form ice crystal, and thus protect cells [20].
The mixed culture was cryopreserved using PEG-2000 as protectant for 6 and 12 months, respectively. The preserved mixed cultures were revived in a water bath, and then were inoculated in the 9 K medium (ferrous and sulphur were added to the culture medium) for 6 d. The growth curves of original and preserved cultures are shown in Figure 1. It can be seen that the cell density of original culture reached the maximum value of 2.43×108 cell/mL in 104 h when using sulphur and ferrous ion as the energy sources. And at the 6th day, the cell density was 2.31×108 cell/mL. However, the preserved culture got relatively low cell density.

Figure 1 Growth curves of original and preserved mixed culture using PEG2000 as cryoprotectant in 9K medium using sulphur and ferrous as energy sources
Particularly, after 12 months of preservation, the cell had a longer lag phase and grew slowly from the beginning. The cell density got the maximum value at the 6th day, with only 1×108 cell/mL. When the culture was preserved for 6 months, the cell density decreased slightly and it was 2.2×108 cell/mL at the 6th day.
After preservation for 6 and 12 months, the cell viability of mixed culture was 87% and 41%, respectively. Furthermore, it was found that the viability of strains using 30% glycerol as protectant was very lower than using PEG-2000 (which was 46% and 29%, respectively). These indicated that PEG-2000 was not only suitable for the storage of pure bioleaching strain [14], but also feasible for the mixed bioleaching culture. For further understanding which species could tolerate the cryopreservation and had the highest survival rate, RFLP technology was used to detect the community structure of the mixed bioleaching culture.
3.2 Bioleaching of chalcopyrite concentrate by original and preserved mixed culture
The mixed culture was enriched from Dexing Copper Mine, China. After several times of adaptation to high density of chalcopyrite, the mixed culture showed good performance for copper extraction (Figure 2). In the bioleaching system,the maximum biomass of the original culture achieved to 1.57×109 cell/mL at the 20th day and the copper extraction rate reached 95.7% at the 25th day. The chalcopyrite bioleaching experiments by the preserved culture were carried out. The results showed that after 6 months of preservation, the biomass of mixed culture reached the maximum value of 1.25×109 cell/mL at the 25th day and the copper extraction rate decreased slightly (94.9%). Furthermore, after 12 months of preservation, the cell growth of mixed culture underwent a relatively longer time of lag phase, and reached the maximum value (9.3×108 cell/mL) at the 30th day. The delay of cell growth could extend the bioleaching time, and the maximum copper extraction rate (93.6%) was obtained at the 35th day.
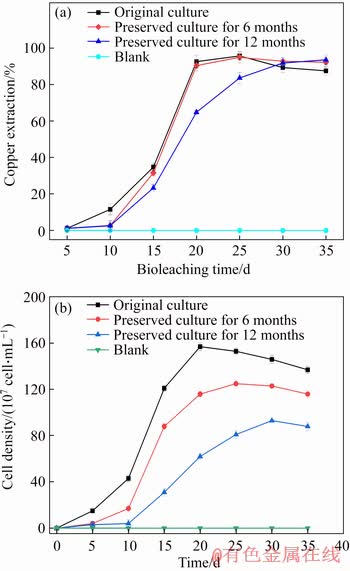
Figure 2 Copper extraction in chalcopyrite bioleaching experiments with original and preserved culture at 20 g/L chalcopyrite density (a) and growth curves of original and preserved mixed culture in chalcopyrite bioleaching experiments (b)
The copper extraction by the preserved culture for 12 months decreased in a different scale compared with original culture. This may be because of the declining cell viability and maximal biomass. On the other hand, preservation would cause some damage to cell activity [21], which maybe affect the ferrous ion or sulphur oxidizing ability during bioleaching. However, after a relatively long time of growth, both the mixed cultures preserved for 6 and 12 months could recover their ability and finally reach relatively high copper extraction. XIAO et al [22] have used an artificial microbial community with four known moderately thermophilic acidophiles (A. caldus S1, S. thermosulfdooxidans ST, L. ferriphilum YSK, and F. thermophilum L1) to explore copper extraction efficiency during chalcopyrite bioleaching, and the results showed that copper extraction efficiency reached the maximum value (about 36%) by the day 24. MA et al [23] studied that the final copper extraction efficiencies of four co-culture microbial groups (A. ferrooxidans, S. thermosulfidooxidans, F. acidiphilum and A. thiooxidans) at 40 °C were 76.34%, 74.84%, 73.57% and 78.37%, respectively. Taken together, the mixed culture of original preserved for 6 and 12 months in the present study showed good performance for copper extraction (about 94%).
3.3 Variation of community structure after cryopreservation for 6 months and 12 months
RFLP technology has been used to reveal the structure of microbial community in Refs. [24-26]. In the 16S rDNA sequences clone library of the original mixed culture, 69 positive clones were re-amplified and then digested, and the banding patterns were grouped into an operational taxonomic unit (OTU) based on the DNA banding pattern of individual clones. A total of 11 (A0_1to A0_11) OTUs were found from the original mixed culture. Phylogenetic tree based on 16S rDNA sequences alignment from original culture is shown in Figure 3. It can be seen that four bacteria species in the original culture were identified as A. ferrooxidans, A. caldus, S. thermotolerans and P. aeruginosa, respectively. Most of these bacteria were always observed in AMD and bioleaching systems [21]. A. ferrooxidans and S. thermotolerans were considered the sulfur-iron oxidizing bacteria. A. caldus was the obligative bacteria for oxidizing sulfur. P. aeruginosa was the heterotrophic bacteria, which can degrade the organic compounds produced by the autotrophic bacteria. The interaction of these microorganisms plays an important role in improving the copper extraction during chalcopyrite bioleaching.
Through calculating the numbers of 16S rDNA clones affiliated to each species, the proportion of each species in the original mixed culture was obtained as shown in Figure 4(a). It is shown that A. ferrooxidans and A. caldus were the dominant bacteria, and their proportions were 44.93% and 31.88%, respectively; while the proportion of S. thermotolerans and P. aeruginosa was 20.29% and 2.9%, respectively. Similar results that A. ferrooxidans and A. caldus were the dominant species at all stages in chalcopyrite bioleaching systems were also reported by XIAO et al [22] and ZHOU et al [27]. When the mixed culture was preserved for 6 months, 70 positive clones were re-amplified and digested, and 11 OTUs were found. The sequencing results showed these four bacteria all existed. This indicated that community structure of the mixed culture after cryopreservation of 6 months didn’t change much (Figure 4(b)), and thus led to the high cell viability. However, when the mixed culture was preserved for 12 months, the community structure has changed largely (Figure 4(c)). We re-amplified and digested 16S rDNA of 67 positive clones, but only 9 OTUs were found. P. aeruginosa was even not detected, and S. thermotolerans decreased largely to only 1.49%. It is shown that the relatively long time preservation like 12 months would obviously lead to the change of community structure. According to the RFLP analysis results, A. ferrooxidans and A. caldus were the predominant microorganisms even after preservation for 12 months. It has been suggested that A. caldus in leachate might indirectly contribute to sulfide mineral oxidation by generating acidity [28]. The extracellular polymeric substances (EPS) and external environmental pressure are important for the function roles of microbial consortium [29-31]. Characterization of the compositions and properties of the EPS of the consortium recovered from the culture after cryopreservation will provide novel insights into the conserved high bioleaching rate of chalcopyrite in future.
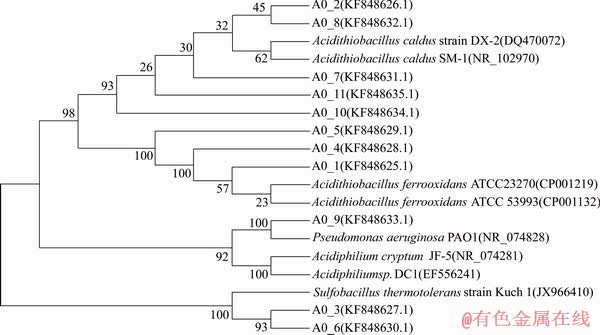
Figure 3 Phylogenetic tree based on 16S rDNA sequences alignment from original culture and related sequences obtained from the NCBI database (The tree was reconstructed using the neighbour-joining method. Numbers at nodes indicated bootstrap values, expressed as percentages of 1000 replications, only values >50% are shown)
4 Conclusions
The mixed culture enriched from Dexing Copper Mine was cryopreserved with PEG-2000 in liquid nitrogen. After cryopreservation for 6 months and 12 months, the maximum copper extraction rate of the mixed culture remained basically stable (about 94%). Additionally, the community composition changed, but the predominant microorganisms still existed. The mixed culture showed good performance for copper extraction. PEG-2000 is a suitable cryoprotectant for the preservation of mixed bioleaching culture. In the near future, longer-term preservation of bioleaching microorganisms for 24 or 36 months will be carried out. The insights gained in this study provide useful guidance to preserve high-efficient mixed bioleaching culture in industrial applications.

Figure 4 Proportion of 16S rDNA sequences copy numbers determined by RFLP in total DNA from original mixed culture (a), 6-month preserved culture (b) and 12-month preserved mixed culture(c)
References
[1] PENG Tang-jian, SHI Li-juan, YU Run-lan, GU Guo-hua, ZHOU Dan, CHEN Miao, QIU Guan-zhou, ZENG Wei-min. Effects of processing pH stimulation on cooperative bioleaching of chalcopyrite concentrate by free and attached cells [J]. Transactions of Nonferrous Metals Society of China, 2016, 26(8): 2220-2229. DOI: 10.1016/S1003-6326(16) 64338-8.
[2] AI Chen-bing, YAN Zhang, CHAI Hong-sheng, GU Tian-yuan, WANG Jun-jun, CHAI Li-yuan, QIU Guan-zhou, ZENG Wei-min. Increased chalcopyrite bioleaching capabilities of extremely thermoacidophilic Metallosphaera sedula inocula by mixotrophic propagation [J]. Journal of Industrial Microbiology & Biotechnology, 2019, 46(8): 1113-1127. DOI: 10.1007/s10295-019-02193-3.
[3] D’HUGUES P, FOUCHER S, GALLE-CAVALLONI P, MORIN D. Continuous bioleaching of chalcopyrite using a novel extremely thermophilic mixed culture [J]. International Journal of Mineral Processing, 2002, 66(1-4): 107-119. DOI: 10.1016/S0301-7516(02)00004-2.
[4] QIU Mu-qing, XIONG Shui-ying, ZHANG Wei-min. Efficacy of chalcopyrite bioleaching using a pure and a mixed bacterium [J]. International Journal of Minerals Metallurgy & Materials, 2006, 13(1): 7-10. DOI: CNKI: SUN:BJKY.0.2006-01-002.
[5] LI Shou-peng, GUO Ning, WU Hai-yan, QIU Guan-zhou, LIU Xin-xing. High efficient mixed culture screening and selected microbial community shift for bioleaching process [J]. Transactions of Nonferrous Metals Society of China, 2011, 21(6): 1383-1387. DOI: 10.1016/s1003-6326(11) 60870-4.
[6] CHEN Yu-zhen, SHAO Kun, GUAN Bing, HOU Pei-bin, ZHANG Hua-ning, BI Zhen-wang. An analysis on the serotypes and antibiotic sensitivities of foodborne Salmonella in Shandong province from 2003 to 2010 [J]. Chinese Journal of Food Hygiene, 2012, 24(1): 9-13. (in Chinese)
[7] RAMANATHAN T, TING Y P. Selective copper bioleaching by pure and mixed cultures of alkaliphilic bacteria isolated from a fly ash landfill site [J]. Water Air & Soil Pollution, 2015, 226(11): 374. DOI: 10.1007/s11270- 015-2641-x.
[8] NIE Yi-lei, CHEN Hong, LUO Li-jin, JIA Wei, CHEN Xing-wei. Screening and identification of mixed culture,and its bioleaching capacity [J]. Biotechnology Bulletin, 2016, 32(8): 177-183. (in Chinese)
[9] DUAN Hong. Preservation and genetic diversity of three Acidithiobacillus caldus strains [D]. Changsha: Central South University, 2013. (in Chinese)
[10] ZHAO Guo, ZHANG Guan. Effect of protective agents, freezing temperature, rehydration media on viability of malolactic bacteria subjected to freeze-drying [J]. Journal of Applied Microbiology, 2010, 99(2): 333-338. DOI: 10.1111/ j.1365-2672.2005.02587.x.
[11] KONEV I E, ZHILINA Z A, CHAMIN N N. Some characteristics of using polyalcohols as cryoprotectors in preserving Actinomyces noursei LIA-0471 [J]. Antibiotiki, 1975, 20(4): 342-345. http://libdb.csu.edu.cn/rwt/SCI/http/ MFZHA63PP7TXE55GNNYG875MMWTGP3JPMNYXN/full_record.do?product=UA&search_mode=GeneralSearch&qid=5&SID=6FRVRcZixS7i7Q1mOIx&page=1&doc=1.
[12] HUBALEK Z. Protectants used in the cryopreservation of microorganisms [J]. Cryobiology, 2003, 46(3): 205-229. DOI: 10.1016/s0011-2240(03)00046-4.
[13] KUWANO K, ARUGA Y, SAGA N. Cryopreservation of clonal gametophytic thalli of Porphyra (Rhodophyta) [J]. Plant Science, 1996, 116(1): 117-124. DOI: 10.1016/0168- 9452(96) 04380-4.
[14] LIU Li-li. Liquid nitrogen preservation and comparative study of molecular polymorphisms of Acidithiobacillus ferrooxidans strains [D]. Changsha: Central South University, 2014. (in Chinese)
[15] WU Xue-ling, HU Qi, HOU Dong-mei, XIN Xiao-hong, MIAO Bo, WANG Yang-yang, LIU Xue-duan, SHEN Li. Preservation efficiency of new cryoprotectant used for Acidithiobacillus ferrooxidans in liquid nitrogen [J]. Transactions of Nonferrous Metals Society of China, 2013, 23(3): 818-823. DOI: 10.1016/S1003-6326(13)62534-0.
[16] WU Xue-ling, XIN Xiao-hong, JIANG Ying, LIANG Ren-xing, YUAN Peng, FANG Chen-xiang. Liquid-nitrogen cryopreservation of three kinds of autotrophic bioleaching bacteria [J]. Transactions of Nonferrous Metals Society of China, 2008, 18(6): 1386-1391. DOI: 10.1016/s1003-6326 (09)60013-3.
[17] ZENG Wei-min, ZHOU Hong-bo, WAN Min-xi, CHAO Wei-liang, XU Ai-ling, LIU Xue-duan, QIU Guan-zhou. Preservation of Acidithiobacillus caldus: A moderately thermophilic bacterium and the effect on subsequent bioleaching of chalcopyrite [J]. Hydrometallurgy, 2009, 96(4): 333-336. DOI: 10.1016/j.hydromet.2008.11.003.
[18] FU Ben-zhong, CHEN Qian-qian, WEI Mi, ZHU Jie-qian, YANG Xin-he, LI Guo-yuan, ZOU Li-ping, WANG Dian-bei. Investigation of walnut bacterial blight pathogens based on 16S-rDNA sequences and RFLP [J]. Journal of Agricultural University of Hebei, 2016, 39(5): 64-68. DOI: 10.13320/j.cnki.jauh2016.0109. (in Chinese)
[19] POSTGATE J R, HUNTER J R. On the survival of frozen bacteria [J]. Journal of General Microbiology, 1961, 26(3): 367-378. DOI: 10.1099/00221287-26-3-367.
[20] ASHWOOD-SMITH M J, WARBY C. Studies on the molecular weight and cryoprotective properties of polyvinylpyrrolidone and dextran with bacteria and erythrocytes [J]. Cryobiology, 1971, 8(5): 453-464. DOI: 10.1016/0011-2240(71)90036-8.
[21] RAWLINGS D E, JOHNSON D B. The microbiology of biomining: Development and optimization of mineral- oxidizing microbial consortia [J]. Microbiology, 2007, 153(2): 315-324. DOI: 10.1099/mic.0.2006/001206-0.
[22] XIAO Yun-hua, LIU Xue-duan, DONG Wei-ling, LIANG Yi-li, NIU Jiao-jiao, GU Ya-bing, MA Li-yuan, HAO Xiao-dong, ZHANG Xian, XU Zhen, YIN Hua-qun. Effects of pyrite and sphalerite on population compositions, dynamics and copper extraction efficiency in chalcopyrite bioleaching process [J]. Archives of Microbiology, 2017, 199(5): 1-10. DOI: 10.1007/s00203-017-1342-9.
[23] MA Li-yuan, WANG Xing-jie, FENG Xue, LIANG Yi-li, XIAO Yun-hua, HAO Xiao-dong, YIN Hua-qun, LIU Hong-wei, LIU Xue-duan. Co-culture microorganisms with different initial proportions reveal the mechanism of chalcopyrite bioleaching coupling with microbial community succession [J]. Bioresource Technology, 2017, 223: 121-130. DOI: 10.1016/j.biortech.2016.10.056.
[24] BATTAGLIA-BRUNET F, CLARENS M, D'HUGUES P, GODON J, FOUCHER S, MORIN D. Monitoring of a pyrite-oxidising bacterial population using DNA single-strand conformation polymorphism and microscopic techniques [J]. Applied Microbiology & Biotechnology, 2002, 60(1, 2): 206-211. DOI: 10.1007/s00253-002-1095-4.
[25] DEMERGASSO C S, GALLEGUILLOS P, ESCUDERO G, ZEPEDA A, DANNY C, CASAMAYOR E O. Molecular characterization of microbial populations in a low-grade copper ore bioleaching test heap [J]. Hydrometallurgy, 2005, 80(4): 241-253. DOI: 10.1016/j.hydromet.2005.07.013.
[26] DEW D W, BUUREN C V, MCEWAN K, BOWKER C. Bioleaching of base metal sulphide concentrates: A comparison of high and low temperature bioleaching [J]. Journal-South African Institute of Mining and Metallurgy, 2000, 100(7): 409-413. http://libdb.csu.edu.cn/rwt/SCI/http/ MFZHA63PP7TXE55GNNYG875MMWTGP3JPMNYXN/full_record.do?product=UA&search_mode=GeneralSearch&qid=1&SID=6FRVRcZixS7i7Q1mOIx&page=1&doc=2.
[27] ZHOU Zhi-jun, YIN Hua-qun, LIU Yi, XIE Ming, QIU Guan-zhou, LIU Xue-duan. Diversity of microbial community at acid mine drainages from Dachang metals-rich mine, China [J]. Transactions of Nonferrous Metals Society of China, 2010, 20(6): 1097-1103. DOI: 10.1016/s1003- 6326(09)60263-6.
[28] TUPIKINA O V, MINNAAR S H, HILLE R P V, WYK N V, RAUTENBACH G F, DEW D, HARRISON S T L. Determining the effect of acid stress on the persistence and growth of thermophilic microbial species after mesophilic colonisation of low grade ore in a heap leach environment [J]. Minerals Engineering, 2013, 53(11): 152-159. DOI: 10.1016/j.mineng.2013.07.015.
[29] AI Chen-bing, YAN Zhang, ZHOU Han, HOU Shan-shan, CHAI Li-yuan, QIU Guan-zhou, ZENG Wei-min. Metagenomic insights into the effects of seasonal temperature variation on functional potentials of activated sludge [J]. Microorganisms, 2019, 7: 713. DOI: 10.3390/ microorganisms7120713.
[30] GAO Qi-yu, TANG De-ping, SONG Peng, ZHOU Jian-ping, LI Hong-yu. Characterization of acylated homoserine lactone derivatives and their influence on biofilms of Acidithiobacillus ferrooxidans BY-3 under arsenic stress [J]. Journal of Central South University, 2020, 27(1): 52-63. DOI: 10.1007/s11771-020-4277-2.
[31] PENG Tang-jian, ZHOU Dan, LIU Ya-nan, YU Run-lan, QIU Guan-zhou, ZENG Wei-min. Effects of pH value on the expression of key iron/sulfur oxidation genes during bioleaching of chalcopyrite on thermophilic condition [J]. Ann Microbiol, 2019, 69(6): 627-635. DOI: 10.1007/ s13213-019-01453-y.
(Edited by YANG Hua)
中文导读
PEG-2000液氮保藏混合浸矿菌对黄铜矿生物浸出的影响
摘要:首先,富集到具有浸矿能力的混合培养物,然后,以PEG-2000作为保护剂,对混合培养物液氮冷冻保藏6个月和12个月,对其保藏前、后黄铜矿的浸出能力、活性和多样性进行比较。结果表明,当用原始培养物浸出黄铜矿时,20天内铜的浸出率为95.7%。冷冻保藏6个月后,混合培养物的铜浸出率在25天内为94.9%,保藏12个月后,在35天内为93.6%。保藏6个月和12个月后,混合菌种的活性分别达到87%和41%。此外,对原始培养物进行生态学分析,鉴定得到Acidithiobacillus ferrooxidans, Acidithiobacillus caldus, Sulfobacillus thermotolerans和Pseudomonas aeruginosa。冷冻保藏12个月后,混合培养物的群落组成发生了变化,但优势菌群依然存在。
关键词:混合培养物;黄铜矿浸出;PEG-2000;冷冻保藏;微生物群落
Foundation item: Projects(31470230, 51320105006, 51604308) supported by the National Natural Science Foundation of China; Project(2017RS3003) supported by the Youth Talent Foundation of Hunan Province of China; Project(2018JJ2486) supported by the Natural Science Foundation of Hunan Province of China; Project(2018WK2012) supported by the Key Research and Development Projects in Hunan Province, China; Project(2018zzts767) supported by the Fundamental Research Funds for the Central Universities, China
Received date: 2019-05-17; Accepted date: 2020-04-13
Corresponding author: ZENG Wei-min, PhD, Professor; Tel: +86-137872885942; E-mail: zengweimin1024@126.com; ORCID: 0000- 0002-7155-6386