
High efficient mixed culture screening and
selected microbial community shift for bioleaching process
LI Shou-peng1, 2, GUO Ning1, 2, WU Hai-yan1, 2,QIU Guan-zhou1, 2, LIU Xin-xing1, 2
1. School of Minerals Processing and Bioengineering, Central South University, Changsha 410083, China;
2. Key Laboratory of Biometallurgy of Ministry of Education, Central South University, Changsha 410083, China
Received 12 June 2010; accepted 24 January 2011
Abstract: To screen the high efficient mixed culture and understand the bioleaching behaviors of mixed culture for low-grade copper sulfide ore bioleaching, ten mixed cultures were collected and screened from different acid mine drainages obtained from sulfide mines of China. The leaching rate was set as criterion to screen the mixed culture and the metagenomic approach. Community genome array (CGA) was used for analyzing the mixed culture microbial community shift during the bioleaching process. The results indicate that the mixed culture obtained from Yinshan(YS) lead-zinc mine in Dexing of Jiangxi province in China reaches the maximum copper extraction (68.89%) during the one bioleaching period of 24 d. CGA results show that YS culture contains nine kinds of bacteria which are belong to six divisions, and the microbial community structure is changing during the bioleaching process. This provides a good way to accelerate the bioleaching process and reveals the microbial community shift during the bioleaching process.
Key words: bioleaching; high efficient mixed culture; community genome array (CGA); microbial community shift
1 Introduction
Bioleaching has been used commercially and offers many cost advantages over traditional techniques, such as energy-efficiency, environmentally friendliness, cost-saving, economic, pollution-free process and low capital requirements. It is quite suitable for treating low-grade ores, waste ores, small and complicated ore bodies and ores that are hard to be recovered by traditional techniques[1]. A lot of studies were carried out in different copper mines around the world by selected bioleaching strains or mixed cultures from acid mine drainage (AMD)[2-4], and the results indicated that the bioleaching capability of mixed culture is obviously better than pure stains[5-8]. However, no research work has focused on comparing bioleaching of low-grade copper sulfide ore using the mixed cultures to choose the high efficient mixed culture.
The understanding of the microbial community structure in the bioleaching systems is very important for the optimization of microbial community by controlling the operating conditions in bioleaching systems, enhancing the bioleaching rate[9]. The microarray has been developed and evaluated for bacterial detection and microbial community analyses in bioleaching systems by ZHOU[10], which has high density, high sensitivity and high quantitative capability.
In this study, ten mixed cultures of acidophilic microorganisms were collected and screened at 30 °C from different AMDs of sulfide mines of China. By comparison experiments in the shake flask, the copper ion bioleaching capabilities of the ten mixed cultures were studied. By the metagenomic approach, the microarray was used for analyzing the mixed culture microbial community shift during the bioleaching process.
2 Materials and methods
2.1 Site description and microorganisms isolation
The mixed cultures of acidophilic microorganisms were collected from AMD samples of sulfide mines of China. The site locations where the samples were collected are shown in Table 1. The indigenous mixed culture is BC and the others are exotic cultures. The pH and temperature of origin samples were 2-3 and 32-35 °C, respectively.
Table 1 Site of samples collected

The AMD samples were filtered by 0.2 μm-nylon filtration membrane (Biobasic Inc.), then the mixed cultures were collected within 24 h after collection and preservated at -20 °C for use. The ten mixed cultures were cultured in the 9 K culture medium under the same condition, containing 0.5 g/L MgSO4・7H2O, 3.0 g/L (NH4)2SO4, 0.5 g/L K2HPO4, 0.1 g/L KCl and 0.01 g/L Ca(NO3)2. 5%(m/V) Dongguashan copper sulfide ore was replaced by FeSO4 as the energy source. It was incubated in rotary shakers with a speed of 180 r/min at 30 °C. The initial pH of the cultures was adjusted to 2.0 with 2.5 mL 1:1 (V/V) H2SO4. After 30 d, the biomass reached 1×107 cells/mL, then microorganisms were collected by centrifugation at speed of 10 000 r/min for 15 min. The sedimentary cells were washed twice with H2SO4 (pH=2.0), and
Table 3 Copper mineral compositions of Dongguashan copper sulfide ore (mass fraction, %)

Table 4 Mineral compositions of Dongguashan copper sulfide ore (mass fraction, %)

2.3 Bioleaching experiment methods
The bioleaching experiments of high efficient mixed culture screening were carried out in 250 mL- Erlenmeyer flasks that were shaken in an air-conditional shaker. At the begining of each experiment, 100 mL 9K culture medium containing 0.5 g/L MgSO4・7H2O, 3.0 g/L (NH4)2SO4, 0.5 g/L K2HPO4, 0.1 g/L KCl and 0.01 g/L Ca(NO3)2 was prepared. The initial pH, temperature, rotation, pulp density and bacteria load of mixed cultures were 2.0, 30 °C, 180 r/min, 10% (m/V) and 10%(m/m) respectively. Two parallel comparative experiments were designed. In control experiment, pure Acidithiobacillus ferrooxidians was inoculated.
Distilled water was added into the flasks on time every day in order to compensate for evaporation losses. Copper concentrations in solution were analyzed every 4 d by atomic absorption spectrophotometry (AAS). Solid residues were filtered, washed, dried, and finally analyzed by X-ray diffractometry (X-RD). Free bacteria in solution were counted in a Petroff-Hausser chamber under an optical microscope.
2.4 Analysis of community genome array (CGA)
YIN et al[11] have reported that a CGA could measure the microbial community structure and the community succession of the bioleaching system successfully. So this CGA was used for measuring the changes in bioleaching system of the mixed culture with the highest copper bioleaching rate. At the beginning of the bioleaching, the 10th day and the 20th day, the samples were taken. The extraction of nucleic acids of the best mixed culture and purification, the oligonucleotide probe (50-mer) design, the microarray construction, the fluorescent labeling of DNA, the hybridization, and the image processing and data analysis were carried out according to the procedure described by YIN et al[11].
3 Results and discussion
3.1 Comparison of bioleaching experiments
Figure 1 shows the copper extraction rate of Dongguashan copper sulfide ore by ten mixed cultures and pure Acidithiobacillus ferrooxidians strain (A.f).

Fig. 1 Copper extraction rate by ten mixed cultures and pure A.f
The results reveal that ten mixed cultures of microorganisms presented in AMD have different capabilities. The mixed culture BC and the mixed culture YS are found to be more efficient than other groups. The mixed culture YTW collected from the acid water reservoir of Yangtaowu, Dexing copper mine, Jiangxi province, has the minimum bioleaching capability of 47.08%. But it is better than the control experiment by pure Acidithiobacillus ferrooxidians strain (30.94%). Furthermore, YS shows the highest extraction rate of Cu. The highest bioleaching rate is 68.69% by the YS. The mixed culture YS is collected from the second heap leach liquor of Yinshan bioleaching plant, Jiangxi province.
89.12% of the Dongguashan copper sulfide ore is primary copper sulfide. Due to the slow kinetics and the high activation energy of chalcopyrite, leaching of chalcopyrite has encountered limited success. However, 68.69% Cu is leached out by YS. In the first 4 d, the bioleaching rate increases faster than in the last 20 d. The possible reasons are as follows.
Firstly, the ore is a kind of high-iron sulfide copper. The total Fe and sulfur are 32.61% and 18.13%, respectively. The iron-oxidizing bacteria in the mixed culture could oxidize Fe (II) to Fe (III) and utilize electrons in their metabolic processes [12]. Fe (III), a powerful oxidant, is able to chemically oxidize the majority of sulfide minerals [13].
CuFeS2 + 4Fe3+→Cu2+ + 2S + 5Fe2+ (1)
4Fe2+ + 4H+ + O2→4Fe3+ + 2H2O (2)
2S + 3O2 + 2H2O→2SO42- + 4H+ (3)
CuFeS2 + 4H+ +1/2O2→Cu2+ + 2S + Fe2++ 2H2O (4)
Secondly, the relation of the magnetite in a lean magnetite ores to gangue is not close; ball milling not only can improve the mineral liberation degree of copper notably but also can improve the particle size characteristics of the ground particles, enhancing the leaching rate.
Finally, this kind of ore is rich in pyrite (11.2%) and pyrrhotite (17.4%). The effect of galvanic interaction between chalcopyrite and pyrite or pyrrhotite [14] can greatly improve the bioleaching rate of chalcopyrite.
3.2 Results of CGA
Analysis of CGA shows that nine kinds of bacteria presented in the bioleaching system of mixed culture of YS. Table 5 shows the results of mutative microbial community structure and foundations of single roles of YS.
In the bioleaching system, nine kinds of bacteria, Acidithiobacillus ferrooxidans, Acidithiobacillus thiooxidans, Acidithiobacillus caldus, Sulfobacillus spp., Alicyclobacillus spp., Arthrobacter globiformis, Acidisphaera spp., Ferromicrobium spp.S-4 and Leptospirillum spp., belong to six divisions, α-Proteobacteria, β-Proteobacteria, γ-Proteobacteria, Firmicutes, Actinobacteria and Nitrospira [2, 15].
For many years, Acidithiobacillus ferrooxidans has been assumed to be the most important microorganism in the bioleaching of sulfide ores at temperature lower than 40 °C, as a iron-oxidizer and sulfur-oxidizer. Using the analysis of the 16S rDNA amplification products of total DNA, the sulphur-oxidising bacteria, Acidithiobacillus thiooxidans and the iron-oxidising bacteria Leptospirillum ferrooxidans, have been found as the dominant populations in bioleaching tanks. At temperatures higher than 40 °C, the moderately thermophilic Acidithiobacillus caldus was also present [16].
Table 5 Mutative microbial community structure and foundations of single role of YS
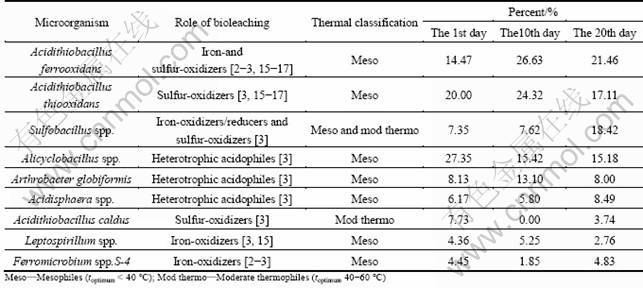
In this study, the dominant bioleaching bacteria were Acidithiobacillus ferrooxidans, Acidithiobacillus thiooxidans, Alicyclobacillus spp. and Sulfobacillus spp.. In the first 10 d of bioleaching, the percentage of Acidithiobacillus ferrooxidans increased from 14.47 % to 26.63%, and it played the most important role. The percentage of three kinds of bacteria, Acidithiobacillus ferrooxidans, Acidithiobacillus thiooxidans, Alicyclobacillus spp., increased from 61.82% to 66.47%. The total biomass increased significantly, too. In the second 10 d, the bioleaching process trended to cease, so the biomass began to decrease, and the percentage of three dominant bacteria decreased to 53.75%. The synergistic effect of autotrophic microorganisms and aerobic-anaerobic microorganisms in the bioleaching system increases the bioleaching rate of the sulfide ore. Alicyclobacillus spp. decreases the toxicity of organic matter accumulation, and Acidithiobacillus thiooxidans avoids the element sulfur membrane of mineral surface, and the copper is leached by Acidithiobacillus ferrooxidans continuously.
4 Conclusions
1) Ten mixed cultures enriched from different places have different capabilities of copper extraction. The mixed cultures BC and YS are more efficient than the others, the mix culture obtained from Yinshan lead-zinc mine (YS) reaches the maximum copper extraction of 68.89% during the one leaching period of 24 d.
2) CGA results show that YS contains nine kinds of bacteria, which belong to six divisions and the microbial community structure is changing during the bioleaching process. The main bioleaching bacteria are Acidithiobacillus ferrooxidans, Acidithiobacillus thiooxidans, Alicyclobacillus spp. and Sulfobacillus spp.. Sulfide-oxidizing bacteria, iron-oxidizing bacteria, the autotrophic bacteria and aerobic-anaerobic bacteria result in synergistic effect of bioleaching process.
References
[1] YIN sheng-hua, WU Ai-xiang, QIU Guan-zhou. Bioleaching of low-grade copper sulphides [J]. Transactions of Nonferrous Metals Society of China, 2008, 18(3): 707-713.
[2] BAKER B J, BANFIELD J F. Microbial communities in acid mine drainage [J]. FEMS Microbiol Ecol, 2003, 44: 139-152.
[3] JOHNSON D B, HALLBERG K B. The microbiology of acidic mine waters [J]. Research in Microbiology, 2003, 154(7): 466-473.
[4] NATARAJAN K A. Microbial aspects of acid mine drainage and its bioremediation [J]. Transactions of Nonferrous Metals Society of China, 2008, 18(6): 1352-1360.
[5] QIU Mu-qing, XIONG Shui-ying, ZHANG Wei-min. Efficacy of chalcopyrite bioleaching using a pure and mixed bacterium [J]. Journal of University of Science and Technology Beijing, 2006, 13(1): 7-10.
[6] DONATI E, CURUTCHET G, POGLIANI C, TEDESCO P. Bioleaching of covellite using pure and mixed cultures of Thiobacillus ferrooxidans and Thiobacillus thiooxidans [J]. Process Biochem, 1996, 31: 129-134.
[7] BEVILAQUA D, LEITE A L L C, O GARCIA, TOUVINEN O H. Oxidation of chalcopyrite by Acidithiobacillus ferroxidans and Acidithiobacillus thiooxidans in shake flasks [J]. Process Biochem, 2002, 38: 587-592.
[8] D’HUGUES P, FOUCHER S, GALLE?-CAVALLONI P, MORIN D. Continuous bioleaching of chalcopyrite using a novel extremely thermophilic mixed culture [J]. Int J Miner Process, 2002, 66: 107-119.
[9] QIU Guan-zhou, LIU Xue-duan, ZHOU Hong-bo. Microbial community structure and function in sulfide ore bioleaching systems [J]. Transactions of Nonferrous Metals Society of China, 2008, 18(6): 1295-1301.
[10] ZHOU Ji-zhong. Microarrays for bacterial detection and microbial community [J]. Current Opinion in Microbiology, 2003, 6: 288-294.
[11] YIN Hua-qun, CAO Lin-hui, QIU Guan-zhou, WANG Dian-zuo, KELLOGG L, ZHOU Ji-zhong, DAI Zhi-min, LIU Xue-duan. Development and evaluation of 50-mer oligonucleotide arrays for detecting microbial populations in acid mine drainages and bioleaching systems [J]. Journal of Microbiological Methods, 2007, 70: 165-178.
[12] THIRD K A, CORD-RUWISCH R, WATLING H R. The role of iron-oxidizing bacteria in stimulation or inhibition of chalcopyrite bioleaching [J]. Hydrometallurgy, 2000, 57: 225-233.
[13] WANG Cui-hong, ZHANG Wei-min. Research progress on promoted leaching of chalcopyrite by iron ion [J]. Hydrometallurgy of China, 2010, 29(2): 67-70. (in Chinese)
[14] ATTIA Y A, EL-ZEKY M. Effects of galvanic interactions of sulfides on extraction of precious metals from refractory complex sulfides by bioleaching [J]. International Journal of Mineral Processing, 1990, 30(1-2): 99-111.
[15] HALLBERG K B. New perspectives in acid mine drainage microbiology [J]. Hydrometallurgy, 2010, 104: 448-453.
[16] FALCO L, POGLIANI C, CURUTCHET G, DONATI E. A comparison of bioleaching of covellite using pure cultures of Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans or a mixed culture of Leptospirillum ferrooxidans and Acidithiobacillus thiooxidans[J]. Hydrometallurgy, 2003, 73: 31-36.
[17] AKCIL A, CIFTCI H, DEVECI H. Role and contribution of pure and mixed cultures of mesophiles in bioleaching of a pyritic chalcopyrite concentrate [J]. Minerals Engineering, 2007, 20: 310-318.
高效浸矿混合菌种的筛选及其
在浸矿过程中的微生物群落结构演变
李寿朋1, 2, 郭 宁1, 2, 武海艳1, 2, 邱冠周1, 2, 刘新星1, 2
1. 中南大学 资源加工与生物工程学院,长沙 410083;
2. 中南大学 生物冶金教育部重点实验室,长沙 410083
摘 要:为了筛选可高效浸出低品位硫化矿的混合菌株,从10个典型的硫化矿矿区的酸性矿坑水中分离富集到混合菌株。以浸出率作为筛选的主要标准,筛选后的高效菌株,利用群落基因组芯片分析混合菌群的组成和过程变化。浸矿持续进行24 d后,采集自江西德兴银山铅锌矿的混合菌浸出率最高,为68.89%。群落基因组芯片结果表明银山菌群包含9种菌,可被分为6类,在浸出过程中群落始终在变化。该研究为加速浸出和了解浸出过程中群落演替提供了一个较好的方法。
关键词:生物浸出;高效混合菌群;群落基因组芯片;微生物群落演替
(Edited by LI Xiang-qun)
Foundation item: Project (50774102) supported by the National Natural Science Foundation of China; Project (2004CB619204) supported by the National Basic Research Program of China
Corresponding author: LIU Xin-xing; Tel: +86-731-88710804; E-mail: xinxingliu@hotmail.com
DOI: 10.1016/S1003-6326(11)60870-4